from phasic import Graph, with_ipv # ALWAYS import phasic first to set jax backend correctly
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
import seaborn as sns
%config InlineBackend.figure_format = 'svg'
from vscodenb import set_vscode_theme, vscode_theme
np.random.seed(42)
set_vscode_theme()
sns.set_palette('tab10')Time inhomogeneity
Phase-type distributions model time-homogeneous Markov jump processes, but phasic has limited support for time-inhomogeneous processs using the two approaches described below:
Step-wise construction
nr_samples = 4
@with_ipv([nr_samples]+[0]*(nr_samples-1))
def coalescent_1param(state):
transitions = []
for i in range(state.size):
for j in range(i, state.size):
same = int(i == j)
if same and state[i] < 2:
continue
if not same and (state[i] < 1 or state[j] < 1):
continue
new = state.copy()
new[i] -= 1
new[j] -= 1
new[i+j+1] += 1
transitions.append([new, [state[i]*(state[j]-same)/(1+same)]])
return transitions
graph = Graph(coalescent_1param)
N = 10000
graph.update_weights([1/N])
graph.plot()PDF/CDF
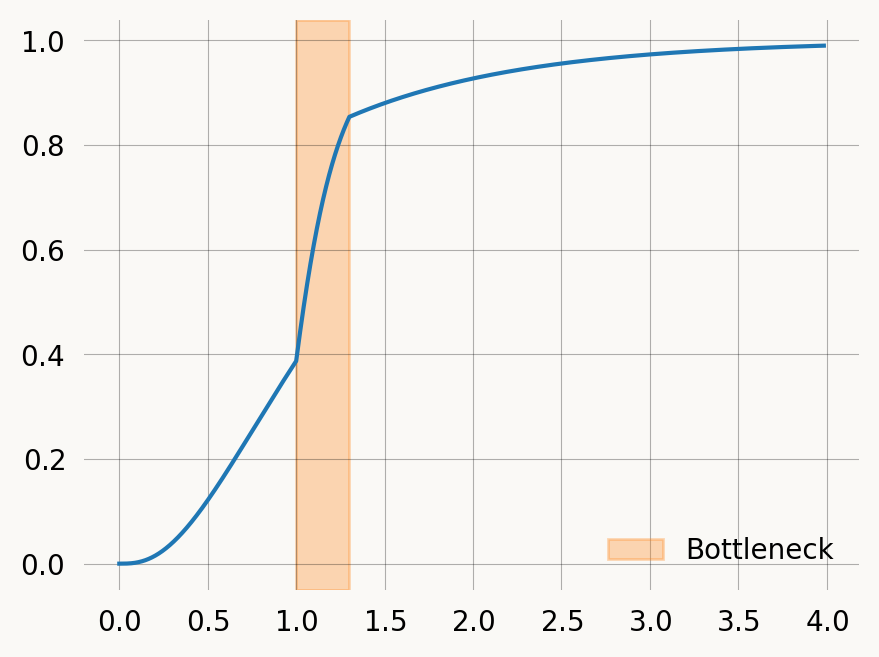
Say we want to model a population bottleneck where N falls to N_{bottle} at time t_{start} and then goes back to N at time t_{end}. To make the approach scale to more changes, and allow more than one parameter we can define changes as a list of (start_time, parameters) tuples:
graph = Graph(coalescent_1param)
N = 1
N_bottle, t_start, t_end = 0.2, 1, 1.3
param_changes = [
(t_start, [1/N_bottle]),
(t_end, [1/N])
]
cdf_cutoff = 0.99
cdf = []
times = []
ctx = graph.distribution_context()
graph.update_weights([1/N])
for change_time, new_params in param_changes:
while ctx.time() < change_time:
cdf.append(ctx.cdf())
times.append(ctx.time())
ctx.step()
if ctx.cdf() >= cdf_cutoff:
break
graph.update_weights(new_params)
while ctx.cdf() < cdf_cutoff:
cdf.append(ctx.cdf())
times.append(ctx.time())
ctx.step()
plt.plot(times, cdf)
plt.axvspan(xmin=t_start, xmax=t_end, alpha=0.3, color='C1', label='Bottleneck')
plt.legend() ;Expectation
If we pick a time far into the future (like 1000), we can integrate under it to find the expectation. accumulated_occupancy computes the time spent in each state up to a time t, so all you have to do is sum these times.
acc_occ = graph.accumulated_occupancy(1000)
acc_occ[0.0,
0.16666666666666563,
0.33333333333332593,
0.3333333333333165,
0.666666666666633,
0.0]np.sum(acc_occ).item(), graph.expectation()(1.499999999999941, 1.4999999999999996)Marginal expectations
We can scale by a reward, and thereby find the marginal expectation of e.g. singleton branch length accumulated by time t:
t = 2
reward_matrix = graph.states().T
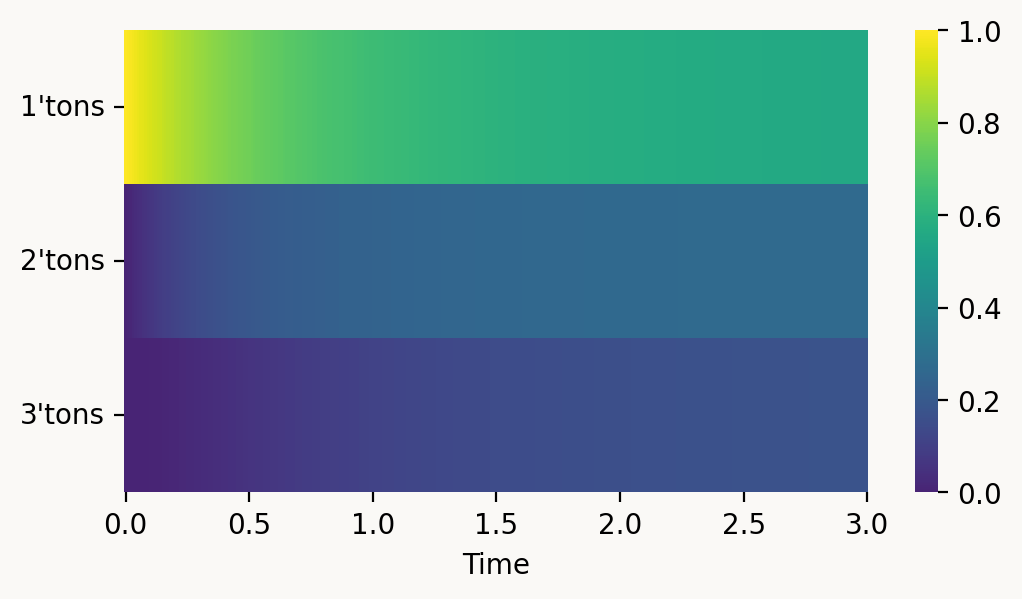
np.sum(graph.accumulated_occupancy(t)*reward_matrix[0]).item()1.8362807228049958That lets us track how the SFS develops over time. Normalized to sum to one, we can see how singletons dominate early in the coalescence process:
@np.vectorize
def brlen_accumulated(i, t):
acc_occ = graph.accumulated_occupancy(t)*reward_matrix[i]
return np.sum(acc_occ).item()
times = np.linspace(0, 3, 301)
tons = list(range(nr_samples-1))
X, Y = np.meshgrid(tons, times, indexing='ij')
result = brlen_accumulated(X, Y)
col_sums = result.sum(axis=0)
result = result / col_sums[np.newaxis, :]
result = pd.DataFrame(
result,
columns=times,
index=[f"{i+1}'tons" for i in range(nr_samples-1)]
)
with vscode_theme(style='ticks'):
fig, ax = plt.subplots(figsize=(6,3))
ax = sns.heatmap(result, cmap='iridis', ax=ax,
xticklabels = 50
)
ax.set_xlabel('Time')
plt.yticks(rotation=0)
plt.show()<Figure size 500x370 with 0 Axes>Moments of epoch-wise time homogeneous phase-type distributions
from phasic import Graph, StateIndexer, PropertySet, Property
import numpy as np
from functools import partial
from itertools import combinations_with_replacement
all_pairs = partial(combinations_with_replacement, r=2)
# nr_samples = 2
# epochs = [0, 1, 2]
# pop_sizes = [1, 5, 10]
# nr_samples = 10
# epochs = [0, 1, 2]
# pop_sizes = [1, 5, 10]
# # epochs = [0, 1, 2, 3, 4, 5]
# # pop_sizes = [1, 5, 10, 2, 4, 1]
# indexer = StateIndexer(
# lineages=[Property('ton', min_value=1, max_value=nr_samples)],
# slots=['epoch']
# )
def coalescent_1param(state, epochs=None, epoch_idx=None, indexer=None):
transitions = []
epoch_idx = int(epoch_idx)
if state[indexer.epoch] != epoch_idx:
return transitions
for i, j in all_pairs(indexer.lineages):
pi = indexer.lineages.index_to_props(i)
pj = indexer.lineages.index_to_props(j)
if state.sum() <= 1:
continue
same = int(pi.ton == pj.ton)
if same and state[i] < 2:
continue
if not same and (state[i] < 1 or state[j] < 1):
continue
new = state.copy()
new[i] -= 1
new[j] -= 1
k = indexer.props_to_index(ton=pi.ton + pj.ton)
new[k] += 1
coeff = np.zeros(len(epochs)+1) # +1 to have a slot for transitions between epochs
coeff[epoch_idx] = state[i]*(state[j]-same)/(1+same)
# print(epoch_idx, coeff)
transitions.append([new, coeff])
# transitions.append([new, [state[i]*(state[j]-same)/(1+same)]])
return transitions
def add_epoch(graph, callback, epochs, epoch_idx, indexer):
epoch = epochs[epoch_idx]
stop_probs = np.array(graph.stop_probability(epoch))
accum_v_time = np.array(graph.accumulated_occupancy(epoch))
with np.errstate(invalid='ignore'):
epoch_trans_rates = stop_probs / accum_v_time
for i in range(1, graph.vertices_length()-1):
if epoch_trans_rates is None or np.isnan(epoch_trans_rates[i]):
continue
if graph.vertex_at(i).edges_length() == 0:
continue
vertex = graph.vertex_at(i)
state = vertex.state()
if not state[indexer.epoch] == epoch_idx - 1:
continue
sister_state = state.copy()
sister_state[indexer.epoch] = epoch_idx
child = graph.find_or_create_vertex(sister_state)
coeff = np.zeros(len(epochs)+1) # +1 to have a slot for transitions between epochs
coeff[-1] = epoch_trans_rates[i]
vertex.add_edge(child, coeff)
graph.extend(callback, epochs=epochs, epoch_idx=epoch_idx, indexer=indexer)
nr_samples = 10
epochs = [0, 1, 2]
pop_sizes = [1, 5, 10]
# epochs = [0, 1, 2, 3, 4, 5]
# pop_sizes = [1, 5, 10, 2, 4, 1]
indexer = StateIndexer(
lineages=[Property('ton', min_value=1, max_value=nr_samples)],
slots=['epoch']
)
ipv = [0] * indexer.state_length
ipv[indexer.props_to_index(ton=1)] = nr_samples
graph = Graph(coalescent_1param,
ipv=ipv,
epochs=epochs,
epoch_idx=0,
indexer=indexer,
)
graph.update_weights([1/size for size in pop_sizes] + [1])
for epoch_idx in range(1, len(epochs)):
graph.update_weights([1/size for size in pop_sizes] + [1])
add_epoch(graph, coalescent_1param, epochs, epoch_idx, indexer)
graph.update_weights([1/size for size in pop_sizes] + [1])
print(graph.moments(5))
graph.plot(size=(12, 8), wrap=False, max_nodes=300)[8.726427931363645, 173.42232048983143, 5233.840746862072, 209923.8992478049, 10506328.513422351]Check that I get the same as Janek for n=10
janek = np.array([8.807791589074768, 177.8449395799212, 5388.12207361224,
216313.46645481227, 10829024.877199283])mine = graph.moments(5)
(mine - janek) / janekarray([-0.00923769, -0.02486784, -0.0286336 , -0.02953846, -0.02979921])SFS
# Get states and remove epoch label column
state_mat = graph.states()
rewards = state_mat[:, :-1] # Remove last column (epoch labels)
# Compute site frequency spectrum
x = np.arange(1, nr_samples)
sfs = np.zeros(nr_samples - 1)
for i in range(nr_samples - 1):
reward_vec = rewards[:, i]
transformed_graph = graph.reward_transform(reward_vec)
sfs[i] = transformed_graph.moments(1)[0]
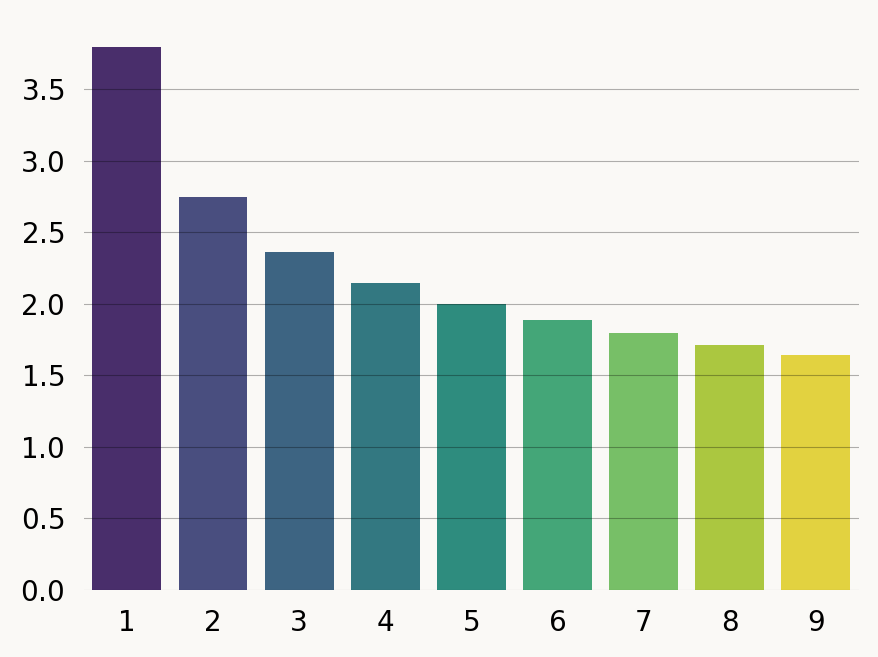
sns.barplot(x=x, y=sfs, hue=x, width=0.8, palette='iridis', legend=False);Comparing to the exact results of Pool and Nielsen for pairwise coelescence time
def exp_coal(g, N):
"""
Compute expected coalescence time in epoch
N is the number of diploid invididuals
g is the number of generations spanned by the epoch
"""
# return 2*N - (g * np.exp(-g/(2*N))) / (1 - np.exp(-g/(2*N)))
return N - (g * np.exp(-g/(N))) / (1 - np.exp(-g/(N)))
def epoch(demog, h, i):
"Recursively compute expected coalescence time across all epoches"
g, N = demog[i]
N *= h
if i == len(demog)-1:
# return 2*N
# return (1-np.exp(-g/(2*N))) * exp_coal(g, N) + np.exp(-g/(2*N)) * (g + epoch(demog, h, i+1))
return N
return (1-np.exp(-g/(N))) * exp_coal(g, N) + np.exp(-g/(N)) * (g + epoch(demog, h, i+1))
def pool_nielsen(gens, Ne, h):
"""
Compute expected coalescence time in units of 2N
Ne is the a list/series of Ne in the epoch.
gens is the a list/series of generation at which an each epoch begins (the last epoch lasts forever)
h is the relative population size, 0.75 for chrX.
"""
epochs = list()
for i in range(len(gens)):
if i == 0:
epochs.append((gens[i+1], Ne[i]))
elif i == len(gens)-1:
epochs.append((None, Ne[i]))
else:
epochs.append((gens[i+1] - gens[i], Ne[i]))
return epoch(epochs, h, 0)n = 10
sampledemog_data = pd.DataFrame(dict(years=[0]+np.logspace(0, 5, n-1, dtype=int, base=10).tolist(),
Ne=np.random.randint(1, 5_000, size=n),
population=['pop']*n
))
sampledemog_data.sort_values('years', inplace=True)
gen_time = 30
exp_coal_time = pool_nielsen(gens=sampledemog_data.years / gen_time,
Ne=sampledemog_data.Ne,
h=1)
# pool_nielsen(gens, Ne, 0.75)
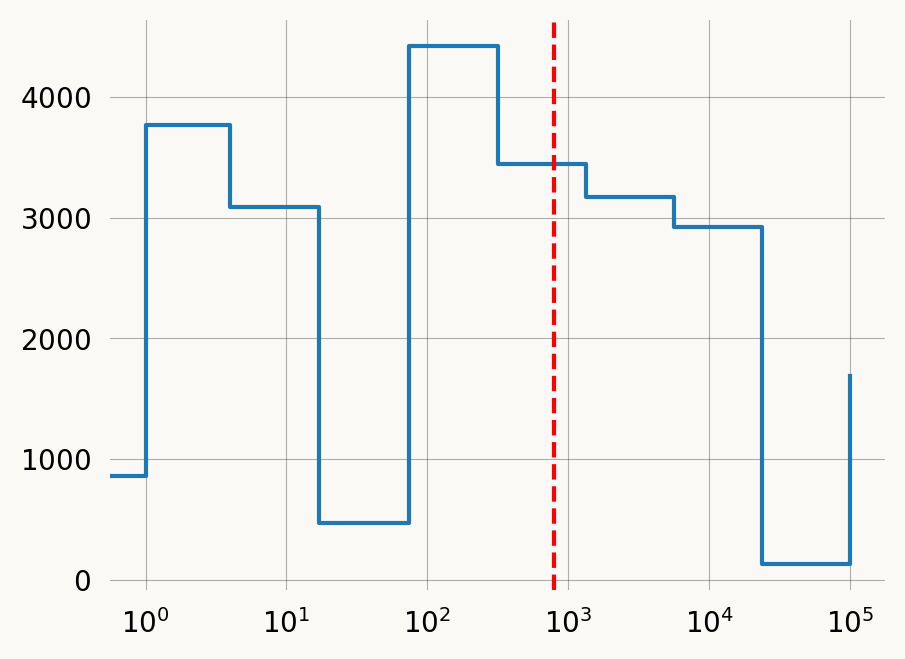
plt.step(sampledemog_data.years, sampledemog_data.Ne, where='post')
plt.gca().set_xscale('log')
plt.gca().axvline(exp_coal_time, color='red', linestyle='--')
plt.show()FIXME: looks like there is a numerical issue here
nr_samples = 2 epochs = [0, 10, 20] pop_sizes = [10, 20, 10]
ipv = [0] * indexer.state_length ipv[indexer.props_to_index(ton=1)] = nr_samples
graph = Graph(coalescent_1param, ipv=ipv, epoch_idx=0, indexer=indexer, )
graph.update_weights([1/size for size in pop_sizes] + [1])
for epoch_idx in range(1, len(epochs)): graph.update_weights([1/size for size in pop_sizes] + [1]) add_epoch(graph, epoch_idx, indexer)
graph.update_weights([1/size for size in pop_sizes] + [1])
graph.expectation(), pool_nielsen(gens=epochs, Ne=pop_sizes, h=1).item()
Discrete distributions
The stepwise technique can be applied to discrete distributions as well in this case the resulting PMF is mathematically exact.
FIXME: update_weights does not work on reward transformed graphs because the reward transformation does not preserve the edges as parameterized
graph = Graph(coalescent_1param, ipv=ipv, epoch_idx=0, indexer=indexer, ) rewards = graph.discretize(lambda state: [0.1 * sum(state)]) rew_graph = graph.reward_transform(rewards)
N = 1 N_bottle, t_start, t_end = 0.2, 1, 1.3
param_changes = [
(t_start, [1/N_bottle]), (t_end, [1/N]) ]
cdf_cutoff = 0.99 cdf = [] times = []
ctx = rew_graph.distribution_context() rew_graph.update_weights([1/N])
for change_time, new_params in param_changes: while ctx.time() < change_time: cdf.append(ctx.cdf()) times.append(ctx.time()) ctx.step() if ctx.cdf() >= cdf_cutoff: break rew_graph.update_weights(new_params) while ctx.cdf() < cdf_cutoff: cdf.append(ctx.cdf()) times.append(ctx.time()) ctx.step()
plt.plot(times, cdf) plt.axvspan(xmin=t_start, xmax=t_end, alpha=0.3, color=‘C1’, label=‘Bottleneck’) plt.legend() ;
Same as the continuous example above, but discrete to account for poisson variance in number of mutations. Notice how the second moment increases:
nr_samples = 10 epochs = [0, 1, 2] pop_sizes = [1, 5, 10]
nr_samples = 2
epochs = [0, 1, 2]
pop_sizes = [1, 2, 10]
indexer = StateIndexer( lineages=[Property(‘ton’, min_value=1, max_value=nr_samples)], slots=[‘epoch’] )
def coalescent_2param(state, epoch_idx=None, indexer=None):
transitions = []
epoch_idx = int(epoch_idx)
if state[indexer.epoch] != epoch_idx:
return transitions
for i, j in all_pairs(indexer.lineages):
pi = indexer.lineages.index_to_props(i)
pj = indexer.lineages.index_to_props(j)
if state.sum() <= 1:
continue
same = int(pi.ton == pj.ton)
if same and state[i] < 2:
continue
if not same and (state[i] < 1 or state[j] < 1):
continue
new = state.copy()
new[i] -= 1
new[j] -= 1
k = indexer.props_to_index(ton=pi.ton + pj.ton)
new[k] += 1
coeff = np.zeros(len(epochs)+1+1) # +1 to have a slot for mutation edges and +1 for edges between epochs
coeff[epoch_idx] = state[i]*(state[j]-same)/(1+same)
# print(epoch_idx, coeff)
transitions.append([new, coeff])
# transitions.append([new, [state[i]*(state[j]-same)/(1+same)]])
return transitionsdef add_epoch(graph, epoch_idx, indexer):
epoch = epochs[epoch_idx]
stop_probs = np.array(graph.stop_probability(epoch))
accum_v_time = np.array(graph.accumulated_occupancy(epoch))
with np.errstate(invalid='ignore'):
epoch_trans_rates = stop_probs / accum_v_time
for i in range(1, graph.vertices_length()-1):
if epoch_trans_rates is None or np.isnan(epoch_trans_rates[i]):
continue
if graph.vertex_at(i).edges_length() == 0:
continue
vertex = graph.vertex_at(i)
state = vertex.state()
if not state[indexer.epoch] == epoch_idx - 1:
continue
sister_state = state.copy()
sister_state[indexer.epoch] = epoch_idx
child = graph.find_or_create_vertex(sister_state)
coeff = np.zeros(len(epochs)+1+1) # +1 to have a slot for transitions between epochs
coeff[-1] = epoch_trans_rates[i]
vertex.add_edge(child, coeff)
graph.extend(coalescent_2param, epoch_idx=epoch_idx, indexer=indexer)ipv = [0] * indexer.state_length ipv[indexer.props_to_index(ton=1)] = nr_samples
graph = Graph(coalescent_2param, ipv=ipv, epoch_idx=0, indexer=indexer, )
def mutation_rate(state): coeff = np.zeros(len(epochs)+1+1) nr_lineages = sum(state[:-2]) coeff[-2] = 1#nr_lineages # TURN THIS BACK TO nr_lineages FOR CORRECT MUTATIONS ON TOTAL BRANCH LENGTH return coeff
graph.update_weights([1/size for size in pop_sizes] + [1] + [1]) # set edge weights
for epoch_idx in range(1, len(epochs)): graph.update_weights([1/size for size in pop_sizes] + [1] + [1]) # set weights of new edges add_epoch(graph, epoch_idx, indexer)
graph.update_weights([1/size for size in pop_sizes] + [1] + [1]) # set weights of new edges rewards = graph.discretize(mutation_rate, skip_existing=True)
print(graph.moments(5, rewards=rewards)) graph.plot(size=(12, 8), max_nodes=300)