from phasic import Graph, with_ipv, StateIndexer, PropertySet, Property
import numpy as np
import seaborn as sns
%config InlineBackend.figure_format = 'svg'
from vscodenb import set_vscode_theme
set_vscode_theme()State properties and indexing
To make it easier to work with complex state spaces phasic sports StateIndexer utility that lets you create the state space from combinations of state properties so you can map between these and the state vector indices. It also provides a flexible way to map between flat the integer indices of the state vector and structured state properties.
This page presents only the core functionality. See the State Indexer Tutorial for more details.
For the coalescent, the state vector indices be referred to by a combination of property values. This is useful when the vector representation of states becomes large. In the coalescent model, there is only a single property (the number of descendants) that maps directly to the state vector index. We need to create a graph more complicated than the standard coalescent to see its use. Lets make a model for the two-locus ancestral recombination graph where we need to keep track of how many descendants a lineage has at each locus.
Basic Property and PropertySet
A Property defines a single dimension of state with a name and valid value range.
# Simple property: number of descendants (0 to 10)
descendants = Property('descendants', max_value=10)
print(f"Property: {descendants.name}")
print(f"Valid range: [{descendants.min_value}, {descendants.max_value}]")
print(f"Number of possible values: {descendants.base}")
print(f"Valid values: {list(descendants)}")Property: descendants
Valid range: [0, 10]
Number of possible values: 11
Valid values: [0, 1, 2, 3, 4, 5, 6, 7, 8, 9, 10]Property with offset min_value: population labels 1-3
population = Property('population', max_value=3, min_value=1)
print(f"Valid range: [{population.min_value}, {population.max_value}]")
print(f"Valid values: {list(population)}")Valid range: [1, 3]
Valid values: [1, 2, 3]Single PropertySet
A StateIndexer with one PropertySet provides a simple state space. Create indexer with single property:
# Create indexer with single property
indexer = StateIndexer(
lineage=[Property('descendants', max_value=10)]
)
print(f"Total states: {indexer.state_length}")
print(f"PropertySet: {indexer.lineage}")Total states: 11
PropertySet: PropertySet('lineage', state_length=11, properties=1)Conversion between index and properties
Convert between flat indices and property values:
# Index to properties (returns dataclass by default)
props = indexer.lineage.i2p(5)
print(f"Index 5: {props}")
print(f"Access via attribute: props.descendants = {props.descendants}")
# Can also get as dict
props_dict = indexer.lineage.i2p(5, as_dict=True)
print(f"As dict: {props_dict}")
# Properties to index
idx = indexer.lineage.p2i({'descendants': 5})
print(f"{{descendants: 5}}: index {idx}")
# Also accepts dataclass
idx = indexer.lineage.p2i(props)
print(f"Dataclass: index {idx}")Index 5: LineageProps(descendants=5)
Access via attribute: props.descendants = 5
As dict: {'descendants': 5}
{descendants: 5}: index 5
Dataclass: index 5Combinatorial State Space with multiple Properties
When you have multiple properties, StateIndexer creates a combinatorial space using mixed-radix encoding.
# Two properties: derived allele status (0 or 1) × descendants (0 to 4)
indexer2 = StateIndexer(
lineage=[
Property('derived', max_value=1), # 2 values: 0, 1
Property('descendants', max_value=4) # 5 values: 0, 1, 2, 3, 4
]
)
print(f"Total states: {indexer2.state_length} (2 × 5 = 10)")
print(f"\nAll states:")
for i in indexer2.lineage:
props = indexer2.lineage.i2p(i)
print(f" Index {i:2d}: derived={props.derived}, descendants={props.descendants}")Total states: 10 (2 × 5 = 10)
All states:
Index 0: derived=0, descendants=0
Index 1: derived=1, descendants=0
Index 2: derived=0, descendants=1
Index 3: derived=1, descendants=1
Index 4: derived=0, descendants=2
Index 5: derived=1, descendants=2
Index 6: derived=0, descendants=3
Index 7: derived=1, descendants=3
Index 8: derived=0, descendants=4
Index 9: derived=1, descendants=4Note: Properties are ordered from least to most significant (like digits in a number). The first property changes fastest when incrementing the index.
Slots for Simple Metadata Storage
“Slots” are single-index storage positions for metadata. Unlike PropertySets, they don’t create combinatorial spaces.
indexer3 = StateIndexer(
'epoch', 'frequency', # Positional slots
lineage=[Property('descendants', max_value=10)]
)
print(f"PropertySet indices: 0-{indexer3.lineage.state_length-1}")
print(f"Slot 'epoch': index {indexer3.epoch}")
print(f"Slot 'frequency': index {indexer3.frequency}")
print(f"Total indices: {indexer3.state_length}")PropertySet indices: 0-10
Slot 'epoch': index 11
Slot 'frequency': index 12
Total indices: 13Index Ranges
Use index_ranges to see how indices are allocated:
print("Index allocation:")
for name, (start, end) in indexer3.index_ranges.items():
if start == end:
print(f" {name:12s}: index {start} (slot)")
else:
print(f" {name:12s}: indices {start}-{end}")Index allocation:
lineage : indices 0-11
epoch : index 11 (slot)
frequency : index 12 (slot)Using Slots with index_to_props
# Check what index 11 (epoch slot) maps to
result = indexer3.i2p(11)
print(f"Index 11:")
print(f" lineage: {result.lineage}")
print(f" epoch: {result.epoch}")
print(f" frequency: {result.frequency}")
# Check a lineage index
result = indexer3.i2p(5)
print(f"\nIndex 5:")
print(f" lineage: {result.lineage}")
print(f" epoch: {result.epoch}")Index 11:
lineage: None
epoch: True
frequency: None
Index 5:
lineage: LineageProps(descendants=5)
epoch: NoneMultiple PropertySets
Organize related properties into separate PropertySets for clarity and modularity.
# Separate lineage properties from metadata
indexer4 = StateIndexer(
'epoch', 'frequency', # Slots
lineage=[
Property('derived', max_value=1),
Property('descendants', max_value=4)
],
metadata=[
Property('time_bin', max_value=100)
]
)
print(f"lineage PropertySet: {indexer4.lineage.state_length} states")
print(f"metadata PropertySet: {indexer4.metadata.state_length} states")
print(f"Total indices: {indexer4.state_length}")
print(f"\nIndex allocation:")
for name, (start, end) in indexer4.index_ranges.items():
if start == end:
print(f" {name:12s}: {start} (slot)")
else:
print(f" {name:12s}: {start}-{end} ({end-start+1} states)")lineage PropertySet: 10 states
metadata PropertySet: 101 states
Total indices: 113
Index allocation:
lineage : 0-10 (11 states)
metadata : 10-111 (102 states)
epoch : 111 (slot)
frequency : 112 (slot)Access PropertySets directly:
print("Lineage states:")
for i in list(indexer4.lineage)[:5]: # First 5
props = indexer4.lineage.i2p(i)
print(f" {i}: {props}")
print("\nMetadata states (first 3):")
for i in list(indexer4.metadata)[:3]:
props = indexer4.metadata.i2p(i)
print(f" {i}: {props}")Lineage states:
0: LineageProps(derived=0, descendants=0)
1: LineageProps(derived=1, descendants=0)
2: LineageProps(derived=0, descendants=1)
3: LineageProps(derived=1, descendants=1)
4: LineageProps(derived=0, descendants=2)
Metadata states (first 3):
0: MetadataProps(time_bin=0)
1: MetadataProps(time_bin=1)
2: MetadataProps(time_bin=2)Flatten Mode - Simplified Access
When working with a single PropertySet, use flatten=True to skip the wrapper object.
# Without flatten (default)
result = indexer4.i2p(5)
print(f"Without flatten: {result}")
print(f" Access: result.lineage.derived = {result.lineage.derived}")
# With flatten=True
props = indexer4.i2p(5, flatten=True)
print(f"\nWith flatten=True: {props}")
print(f" Direct access: props.derived = {props.derived}")Without flatten: IndexResult(lineage=LineageProps(derived=1, descendants=2), metadata=None, epoch=None, frequency=None)
Access: result.lineage.derived = 1
With flatten=True: LineageProps(derived=1, descendants=2)
Direct access: props.derived = 1When iterating over a PropertySet, flatten=True is usually what you want:
# Common pattern: roundtrip within a PropertySet
print("Roundtrip demonstration:")
for i in list(indexer4.lineage)[:5]:
props = indexer4.i2p(i, flatten=True) # Get properties
idx_back = indexer4.p2i(props) # Convert back (auto-detects PropertySet!)
print(f" {i} -> {props} → {idx_back} ->")Roundtrip demonstration:
0 -> LineageProps(derived=0, descendants=0) → 0 ->
1 -> LineageProps(derived=1, descendants=0) → 1 ->
2 -> LineageProps(derived=0, descendants=1) → 2 ->
3 -> LineageProps(derived=1, descendants=1) → 3 ->
4 -> LineageProps(derived=0, descendants=2) → 4 ->Auto-Detection Magic
When property names are unique across PropertySets, StateIndexer automatically detects which PropertySet you’re referring to.
# Properties are unique: 'derived', 'descendants' only in lineage; 'time_bin' only in metadata
indexer5 = StateIndexer(
lineage=[
Property('derived', max_value=1),
Property('descendants', max_value=4)
],
metadata=[
Property('time_bin', max_value=100)
]
)
# Auto-detection works:
idx = indexer5.p2i({'derived': 1, 'descendants': 2})
print(f"Auto-detected 'lineage': {idx}")
idx = indexer5.p2i({'time_bin': 50})
print(f"Auto-detected 'metadata': {idx}")
# Also works with kwargs
idx = indexer5.p2i(derived=1, descendants=2)
print(f"Kwargs also work: {idx}")Auto-detected 'lineage': 5
Auto-detected 'metadata': 60
Kwargs also work: 5When Auto-Detection Fails: Helpful Errors
If property names are ambiguous or not found, you get a helpful error message:
# Create indexer with ambiguous property 'count' in both PropertySets
indexer_ambig = StateIndexer(
lineage=[Property('count', max_value=10)],
metadata=[Property('count', max_value=100)]
)
try:
idx = indexer_ambig.p2i({'count': 5})
except ValueError as e:
print("Error (ambiguous):")
print(e)Error (ambiguous):
Cannot auto-detect PropertySet: property names ['count'] match multiple PropertySets: ['lineage', 'metadata'].
Matching PropertySets:
lineage: ['count']
metadata: ['count']
Use indexer.pset_name.p2i(props) to specify explicitly.Property not found:
try:
idx = indexer5.p2i({'nonexistent': 5})
except ValueError as e:
print("Error (not found):")
print(e)Error (not found):
Cannot auto-detect PropertySet: property names ['nonexistent'] not found in any PropertySet.
Available properties by PropertySet:
lineage: ['derived', 'descendants']
metadata: ['time_bin']
Use indexer.pset_name.p2i(props) instead.Explicit PropertySet Selection
When auto-detection doesn’t work (or you want to be explicit), specify the PropertySet:
# Explicit PropertySet access
idx = indexer_ambig.lineage.p2i({'count': 5})
print(f"lineage.count=5: index {idx}")
idx = indexer_ambig.metadata.p2i({'count': 5})
print(f"metadata.count=5: index {idx}")
# Or via props_to_index with PropertySet name
idx = indexer_ambig.props_to_index('lineage', {'count': 5})
print(f"Explicit 'lineage': index {idx}")lineage.count=5: index 5
metadata.count=5: index 5
Explicit 'lineage': index 5Advanced Features
Query all states matching a partial property specification:
indexer6 = StateIndexer(
lineage=[
Property('derived', max_value=1),
Property('descendants', max_value=4)
]
)
# Find all states with derived=1 (any number of descendants)
indices = indexer6.lineage.p2i({'derived': 1})
print(f"All states with derived=1: {indices}")
print("Properties:")
for i in indices:
print(f" {i}: {indexer6.lineage.i2p(i)}")All states with derived=1: [1 3 5 7 9]
Properties:
1: LineageProps(derived=1, descendants=0)
3: LineageProps(derived=1, descendants=1)
5: LineageProps(derived=1, descendants=2)
7: LineageProps(derived=1, descendants=3)
9: LineageProps(derived=1, descendants=4)# Find all states with descendants=3
indices = indexer6.lineage.p2i({'descendants': 3})
print(f"\nAll states with descendants=3: {indices}")
print("Properties:")
for i in indices:
print(f" {i}: {indexer6.lineage.i2p(i)}")
All states with descendants=3: [6 7]
Properties:
6: LineageProps(derived=0, descendants=3)
7: LineageProps(derived=1, descendants=3)Vectorized Operations
Convert arrays of indices efficiently:
indices = np.array([0, 2, 4, 6, 8])
results = indexer6.lineage.i2p(indices)
print("Vectorized conversion:")
for i, props in zip(indices, results):
print(f" {i}: {props}")Vectorized conversion:
0: LineageProps(derived=0, descendants=0)
2: LineageProps(derived=0, descendants=1)
4: LineageProps(derived=0, descendants=2)
6: LineageProps(derived=0, descendants=3)
8: LineageProps(derived=0, descendants=4)Get as 2D array:
values = indexer6.lineage.i2p(indices, as_values=True)
print(f"\nAs 2D array:\n{values}")
print(f"Shape: {values.shape} (n_indices, n_properties)")
As 2D array:
[[0 0]
[0 1]
[0 2]
[0 3]
[0 4]]
Shape: (5, 2) (n_indices, n_properties)Best Practices
Use descriptive property names:
Keep property names unique across PropertySets
# Good: enables auto-detection
indexer_unique = StateIndexer(
lineage=[Property('descendants', max_value=10)],
metadata=[Property('time_bin', max_value=100)] # Different names!
)
# Now auto-detection works
idx = indexer_unique.p2i({'descendants': 5}) # Auto-detects 'lineage'
print(f"Auto-detection works: {idx}")Auto-detection works: 5Use flatten=True when iterating a single PropertySet:
# Good: clean and direct
for i in indexer_unique.lineage:
props = indexer_unique.i2p(i, flatten=True)
idx_back = indexer_unique.p2i(props) # Auto-detects + accepts dataclass
assert i == idx_back
print("✓ Roundtrip successful for all states")✓ Roundtrip successful for all statesUse PropertySet methods directly when explicit:
# Good: explicit and clear
for i in indexer_unique.lineage:
props = indexer_unique.lineage.i2p(i)
idx_back = indexer_unique.lineage.p2i(props)
assert i == idx_back
print("✓ Explicit PropertySet access")✓ Explicit PropertySet accessnr_samples = 4
indexer = StateIndexer(
n_descendants=[
Property('locus1', max_value=nr_samples),
Property('locus2', max_value=nr_samples)
]
)
indexerStateIndexer(n_descendants=[locus1:0-4, locus2:0-4])indexer.state_length25indexer.n_descendantsPropertySet('n_descendants', state_length=25, properties=2)Each index in the state vector maps to a set of property values:
print('index', 'locus1', 'locus2', sep='\t')
# you can iterate over indices of a property set
for i in indexer.n_descendants:
props = indexer.n_descendants.index_to_props(i)
print(i, props.locus1, props.locus2, sep='\t')index locus1 locus2
0 0 0
1 1 0
2 2 0
3 3 0
4 4 0
5 0 1
6 1 1
7 2 1
8 3 1
9 4 1
10 0 2
11 1 2
12 2 2
13 3 2
14 4 2
15 0 3
16 1 3
17 2 3
18 3 3
19 4 3
20 0 4
21 1 4
22 2 4
23 3 4
24 4 4You can map back from properties to index:
props = indexer.n_descendants.index_to_props(5)
props.locus1, props.locus2(0, 1)indexer.n_descendants.props_to_index(props)5indexer.n_descendants.props_to_index(locus1=props.locus1 + 1,
locus2=props.locus2)6Two-locus ARG:
# zero state vector with appropriate size
initial = [0] * indexer.state_length
# set initial state with all lineages having one descendant at both loci
initial[indexer.props_to_index(locus1=1, locus2=1)] = nr_samples
@with_ipv(initial)
def two_locus_arg(state, indexer=None, N=None, R=None):
transitions = []
if state.sum() <= 1: return transitions
for i in range(indexer.state_length):
if state[i] == 0: continue
props_i = indexer.n_descendants.index_to_props(i)
for j in range(i, indexer.state_length):
if state[j] == 0: continue
props_j = indexer.n_descendants.index_to_props(j)
same = int(i == j)
if same and state[i] < 2:
continue
if not same and (state[i] < 1 or state[j] < 1):
continue
child = state.copy()
child[i] -= 1
child[j] -= 1
locus1 = props_i.locus1 + props_j.locus1
locus2 = props_i.locus2 + props_j.locus2
if locus1 <= nr_samples and locus2 <= nr_samples:
child[indexer.props_to_index(locus1=locus1, locus2=locus2)] += 1
transitions.append([child, state[i]*(state[j]-same)/(1+same)/N])
if state[i] > 0 and props_i.locus1 > 0 and props_i.locus2 > 0:
child = state.copy()
child[i] -= 1
child[indexer.props_to_index(locus1=props_i.locus1, locus2=0)] += 1
child[indexer.props_to_index(locus1=0, locus2=props_i.locus2)] += 1
transitions.append([child, R])
return transitions
graph = Graph(two_locus_arg,
N=1, R=1,
indexer=indexer) # passing indexer as argument
graph.plot(max_nodes=200)# def fun(i):
# i in indexer.props_to_index({'locus1': True})
# graph.plot(max_nodes=200, by_index=fun)graph.expectation()2.0091445466464215reward_matrix = graph.states().TIf you pass only a subset of the state properties, props_to_index will return all the indices matching that property. This lets you get the get the relevant indices for use as rewards. Say, you want the total branch length of lineages with one descendant at locus 1 and any number of descendants at locus 2:
idx_singleton_loc1 = indexer.n_descendants.props_to_index(locus1=1)
idx_singleton_loc1array([ 1, 6, 11, 16, 21])Now we can get those reward vectors and sum them:
reward_matrix = graph.states().T
singleton_rewards_loc1 = reward_matrix[idx_singleton_loc1].sum(axis=0)
singleton_rewards_loc1array([0, 4, 2, 4, 0, 1, 2, 2, 2, 4, 4, 0, 0, 1, 1, 0, 1, 2, 2, 2, 2, 1,
2, 2, 2, 2, 4, 4, 4, 0, 0, 0, 0, 1, 1, 0, 1, 0, 0, 0, 1, 1, 1, 2,
2, 2, 2, 2, 2, 2, 1, 2, 1, 2, 0, 1, 2, 2, 4, 4, 4, 4, 4, 0, 0, 0,
1, 1, 1, 0, 0, 0, 0, 0, 0, 1, 1, 1, 2, 2, 2, 2, 2, 2, 2, 1, 2, 0,
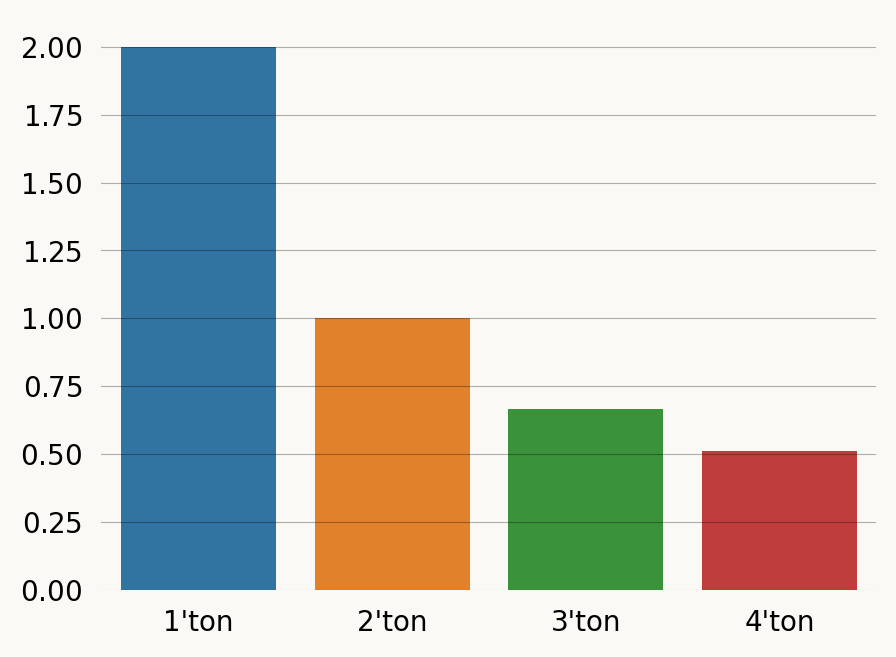
1, 1, 2, 4, 4, 4, 4, 0, 0, 0, 1, 1, 2, 2, 0, 0, 1, 4, 4, 4, 0, 4])graph.expectation(singleton_rewards_loc1)1.9999999999999991If we repeat the exercise for all numbers of descendants at locus 1, we can get the standard SFS (we we should), except that the expected 4’ton branch length is not zero. This is because there can be a 4’ton branch from the TMRCA of the samples at locus 1 and the TMRCA for the full ARG: