Lets try to estimate both coalescent and migration rate in a two population model.
from phasic import (# ALWAYS import phasic first to set jax backend correctly import numpy as npimport jax.numpy as jnpimport pandas as pdfrom typing import Optionalimport matplotlib.pyplot as pltfrom matplotlib.colors import LogNormimport seaborn as sns% config InlineBackend.figure_format = 'svg' from tqdm.auto import tqdmfrom vscodenb import set_vscode_theme42 )'tab10' )'WARNING' )
<phasic.logging_config.set_log_level at 0x15f183e10>
= 2 = StateIndexer(= ['pop1' , min_value= 0 , max_value= nr_samples),'pop2' , min_value= 0 , max_value= nr_samples),'in_pop' , min_value= 1 , max_value= 2 ),= [0 ] * indexer.state_length# set initial state with all lineages having one descendant at both loci = 1 , pop2= 0 , in_pop= 1 )] = nr_samples@with_ipv (initial)def two_island(state):= []if state[indexer.descendants.indices()].sum () <= 1 :return transitionsfor i in range (indexer.descendants.state_length):if state[i] == 0 : continue = indexer.descendants.index_to_props(i)for j in range (i, indexer.descendants.state_length):if state[j] == 0 : continue = indexer.descendants.index_to_props(j)if props_j.in_pop != props_i.in_pop:continue = int (i == j)if same and state[i] < 2 :continue if not same and (state[i] < 1 or state[j] < 1 ):continue = state.copy()-= 1 -= 1 = props_i.pop1 + props_j.pop1= props_i.pop2 + props_j.pop2if des_pop1 <= nr_samples and des_pop2 <= nr_samples:= indexer.descendants.props_to_index(= des_pop1, = des_pop2, = props_i.in_pop+= 1 * (state[j]- same)/ (1 + same), 0 ]])if state[i] > 0 := state.copy()= 2 if props_i.in_pop == 1 else 1 = state.copy()-= 1 = indexer.descendants.props_to_index(= props_i.pop1, = props_i.pop2, = other_pop+= 1 0 , state[i]]])return transitions= Graph(two_island) = [0.7 , 0.3 ]def label(state):= sum ([state[i] * bool (indexer.index_to_props(i).descendants.in_pop == 1 ) for i in indexer])= sum ([state[i] * bool (indexer.index_to_props(i).descendants.in_pop == 2 ) for i in indexer])return f"pop1: { nr_pop1} \n pop2: { nr_pop2} " = 'LR' , nodesep= 0.3 , ranksep= 2 ,= 10 , # label_fmt=False, = label)
= 1.2e-4 = graph.joint_prob_graph(indexer, = ['pop1' , 'pop2' ],= 1 ,= 1 , = mutation_rate)= 'LR' , nodesep= 0.3 , ranksep= 2 ,= 10 , label_fmt= False , by_state= label)
= joint_prob_graph.joint_prob_table()
t_vertex_index
18
0
0
0
0
0
0
0.999041
19
0
1
0
0
0
0
0.000479
20
0
0
0
1
0
0
0.000479
If an indexer has a prop set with multiple props and I call When I call props_to_index specifying only a single prop i get:
ValueError Traceback (most recent call last) Cell In[36], line 5 3 values = {p.name: list(range(p.min_value, p.max_value+1)) for p in indexer.descendants.properties} 4 for tup in product(*[values[p] for p in reward_only]): —-> 5 print(indexer.props_to_index(**dict(zip(reward_only, tup))))
File ~/phasic/.pixi/envs/default/lib/python3.11/site-packages/phasic/state_indexing.py:1491, in StateIndexer.props_to_index(self, pset_name, props, kwargs) 1485 raise KeyError( 1486 f”PropertySet ‘{actual_pset_name}’ not found. ” 1487 f”Available PropertySets: {available}” 1488 ) 1490 local_index = self._property_sets[actual_pset_name].props_to_index(actual_props, kwargs) -> 1491 return self._compose_index(actual_pset_name, local_index)
File ~/phasic/.pixi/envs/default/lib/python3.11/site-packages/phasic/state_indexing.py:1217, in StateIndexer._compose_index(self, name, local_index) 1215 raise ValueError(f”PropertySet ‘{name}’ requires local_index parameter”) 1216 pset = self._property_sets[name] -> 1217 if not (0 <= local_index < pset.state_length): 1218 raise IndexError( 1219 f”Local index {local_index} out of range for PropertySet ‘{name}’ ” 1220 f”(valid range: [0, {pset.state_length}))” 1221 ) 1222 return self._offsets[name] + local_index
ValueError: The truth value of an array with more than one element is ambiguous. Use a.any() or a.all()
# nr_samples = 2 # indexer = StateIndexer( # descendants=[ # Property('pop1', min_value=0, max_value=nr_samples), # Property('pop2', min_value=0, max_value=nr_samples), # Property('in_pop', min_value=1, max_value=2), # ]) # initial = [0] * indexer.state_length # # set initial state with all lineages having one descendant at both loci # initial[indexer.descendants.props_to_index(pop1=1, pop2=0, in_pop=1)] = nr_samples # @with_ipv(initial) # def two_island(state): # transitions = [] # if state[indexer.descendants.indices()].sum() <= 1: # return transitions # for i in range(indexer.descendants.state_length): # if state[i] == 0: continue # props_i = indexer.descendants.index_to_props(i) # for j in range(i, indexer.descendants.state_length): # if state[j] == 0: continue # props_j = indexer.descendants.index_to_props(j) # if props_j.in_pop != props_i.in_pop: # continue # same = int(i == j) # if same and state[i] < 2: # continue # if not same and (state[i] < 1 or state[j] < 1): # continue # child = state.copy() # child[i] -= 1 # child[j] -= 1 # des_pop1 = props_i.pop1 + props_j.pop1 # des_pop2 = props_i.pop2 + props_j.pop2 # if des_pop1 <= nr_samples and des_pop2 <= nr_samples: # k = indexer.descendants.props_to_index( # pop1=des_pop1, # pop2=des_pop2, # in_pop=props_i.in_pop # ) # child[k] += 1 # transitions.append([child, [state[i]*(state[j]-same)/(1+same), 0]]) # if state[i] > 0: # child = state.copy() # other_pop = 2 if props_i.in_pop == 1 else 1 # child = state.copy() # child[i] -= 1 # k = indexer.descendants.props_to_index( # pop1=props_i.pop1, # pop2=props_i.pop2, # in_pop=other_pop # ) # child[k] += 1 # transitions.append([child, [0, state[i]]]) # return transitions # graph = Graph(two_island) # true_theta = [0.7, 0.3] # graph.update_weights(true_theta) # graph.plot(rankdir='LR', by_index=lambda i: f"Desc in pop1: {indexer.index_to_props(i).descendants.pop1}")
# graph.plot(rankdir='LR', by_state=lambda s: f"In pop1: {s[indexer.descendants.props_to_index(pop1=1)].sum()}")
= graph.sample(1000 )
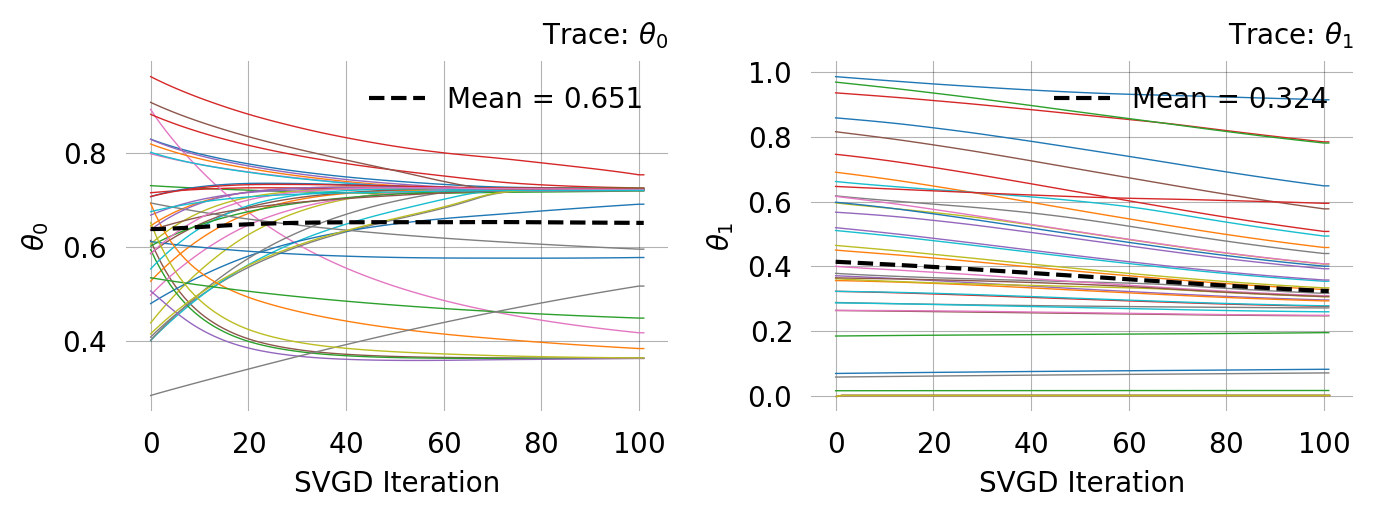
= graph.svgd(observations, # n_particles=40, n_iterations=100
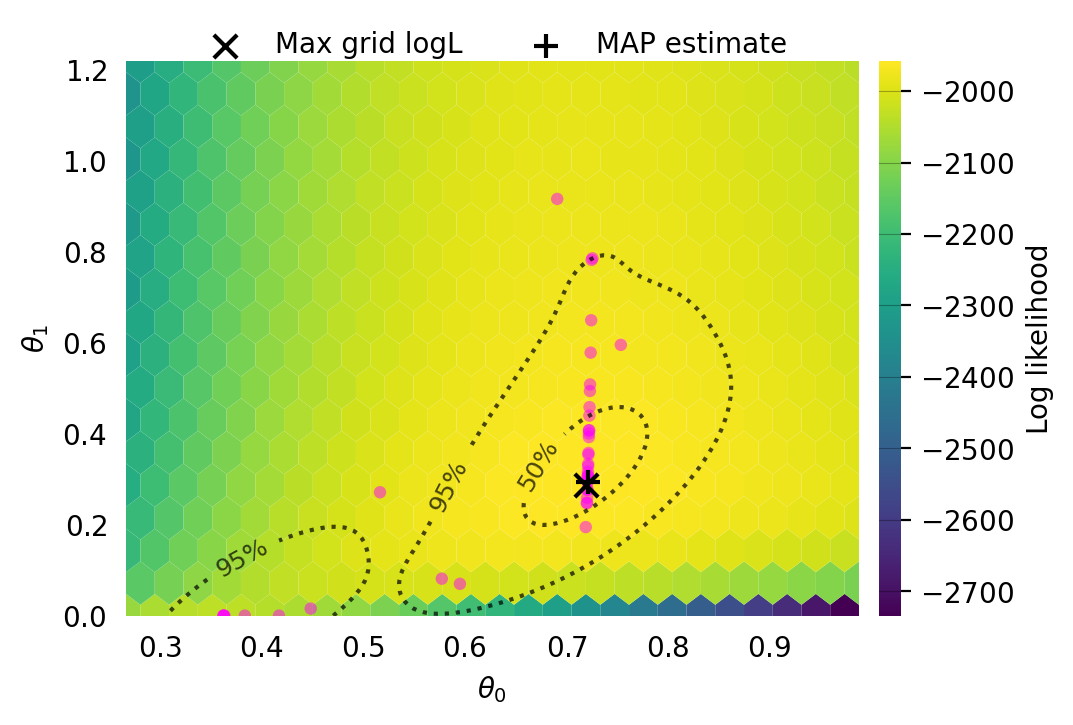
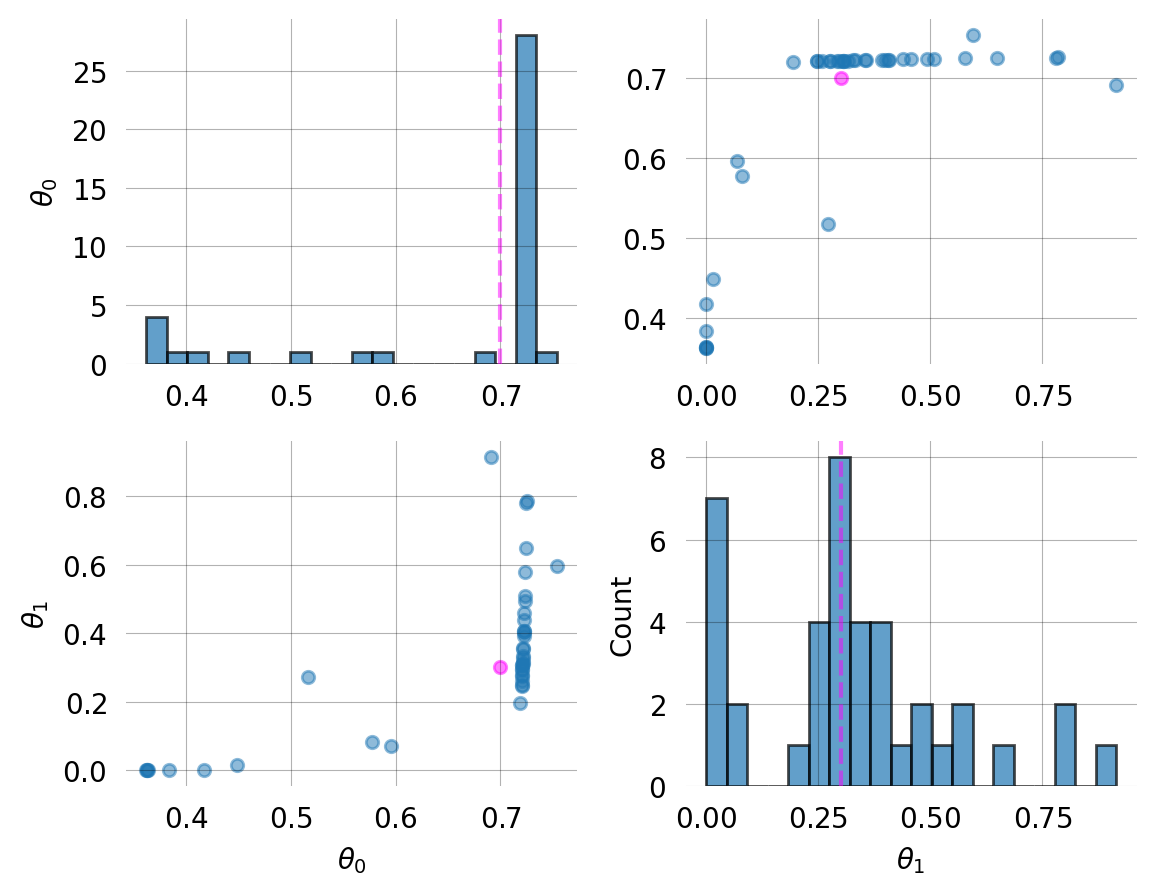
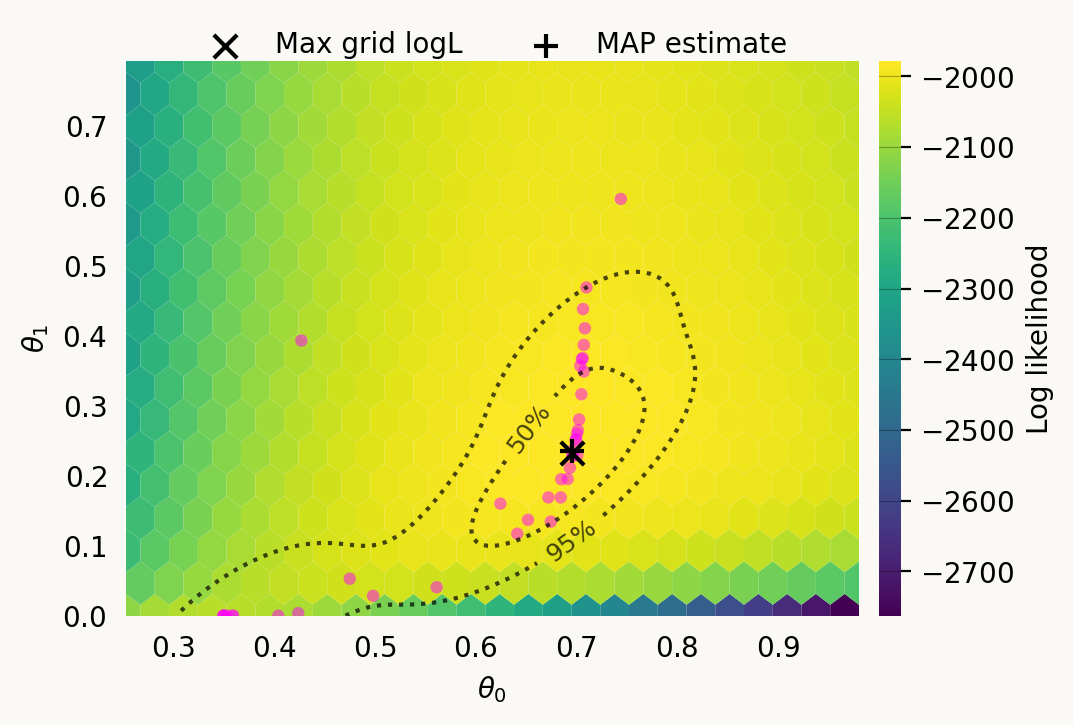
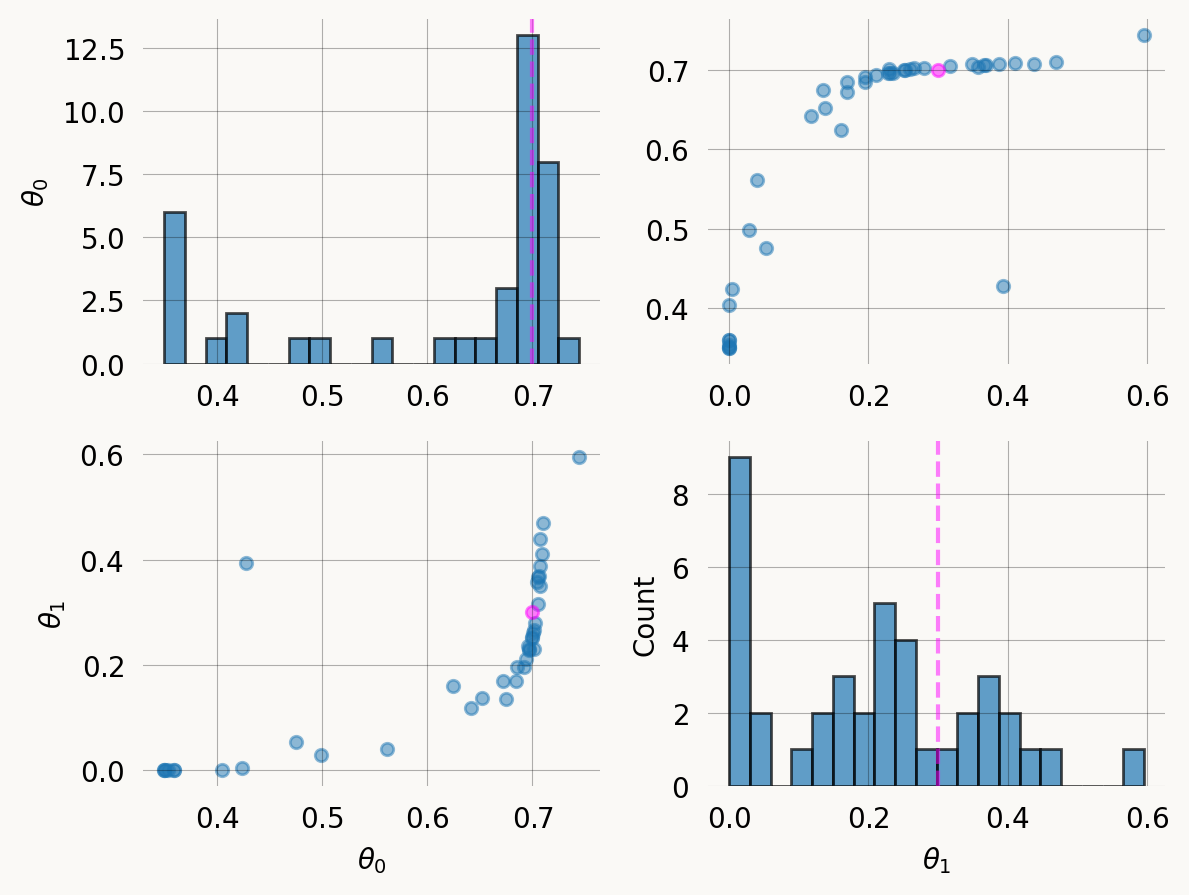
Parameter Fixed MAP Mean SD HPD 95% lo HPD 95% hi
0 No 0.6957 0.6084 0.1367 0.3498 0.7087
1 No 0.2356 0.2075 0.1545 0.0000 0.4384
Particles: 40, Iterations: 100
= true_theta)
#svgd.animate_pairwise(true_theta=true_theta)