from phasic import Graph, with_ipv # ALWAYS import phasic first to set jax backend correctly
import sys
import numpy as np
import pandas as pd
import matplotlib.pyplot as plt
from matplotlib.colors import LogNorm
import seaborn as sns
%config InlineBackend.figure_format = 'svg'
from vscodenb import set_vscode_theme
np.random.seed(42)
set_vscode_theme()
sns.set_palette('tab10')Discrete phase-type distributions
Consider this simple graph representing an exponential distribution. If you must, you can make it the coalescence time of two lineages in very small population:
def exponential(state):
transitions = [] # List for transitions from state
if state[0] < 2: # Control transitions allowed
child = state.copy() # Make a copy to modify
child[0] += 1 # Make the child state
trans = [child, 2.0] # Pair the child with a transition rate
transitions.append(trans) # Append transition
return transitions
graph = Graph(exponential, ipv=[1])
graph.plot()def exponential(rate):
graph = Graph(1)
vertex = graph.find_or_create_vertex([2])
graph.starting_vertex().add_edge(vertex, 1)
abs_vertex = graph.find_or_create_vertex([1])
vertex.add_edge(abs_vertex, rate)
return graph
graph = exponential(0.4)
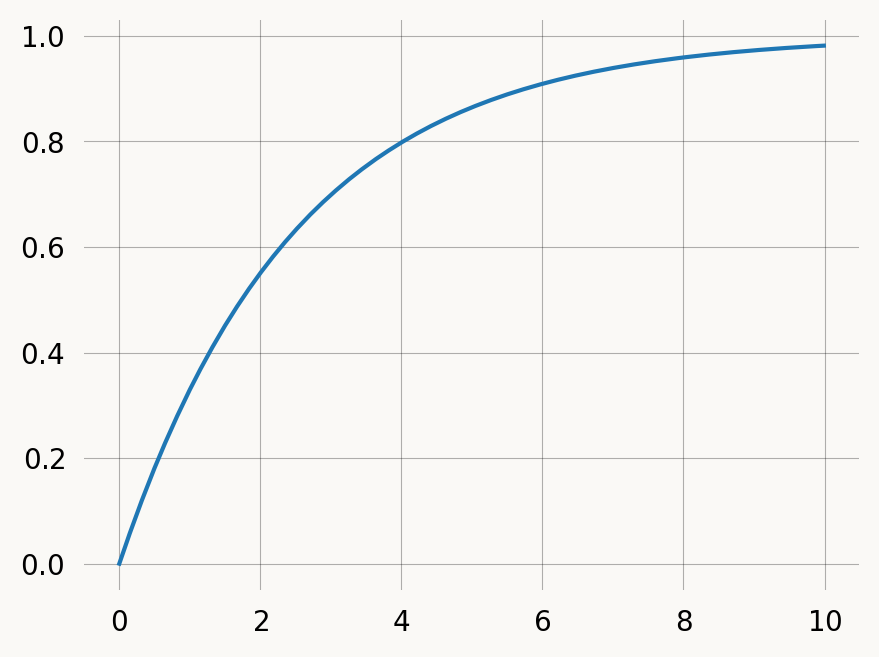
graph.plot()rate = 0.4
x = np.arange(0, 10, 0.01)
graph = exponential(rate)
plt.plot(x, graph.cdf(x))
plt.show()Discrete phase-type distributions represent the the number of jumps in a Markov Chain before absorption. We can discretize our graph it to get a geometric distribution:
The discretize method changes to graph in-place, by connecting each non-absorbing vertex in the graph to an auxiliary vertex with a specified rate. Each AUX vertex connects to its parent with rate 1. Once AUX vertices are added, the graph is normalized so the sum of edge weights sum to 1 for each vertex. discretize returns the graph indices of the added AUX vertices.
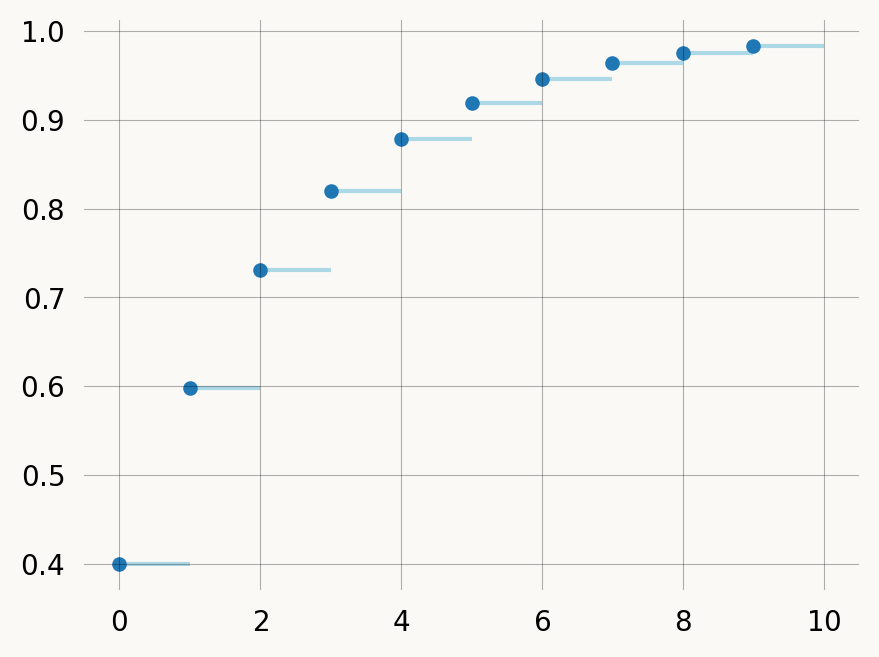
x = np.arange(10)
# cdf = graph.reward_transform(rewards).cdf(x, discrete=True)
cdf = graph.reward_transform(rewards).cdf(x)
plt.hlines(y=cdf, xmin=x, xmax=x+1, zorder=1, color='lightblue')
plt.scatter(x, cdf, s=18)
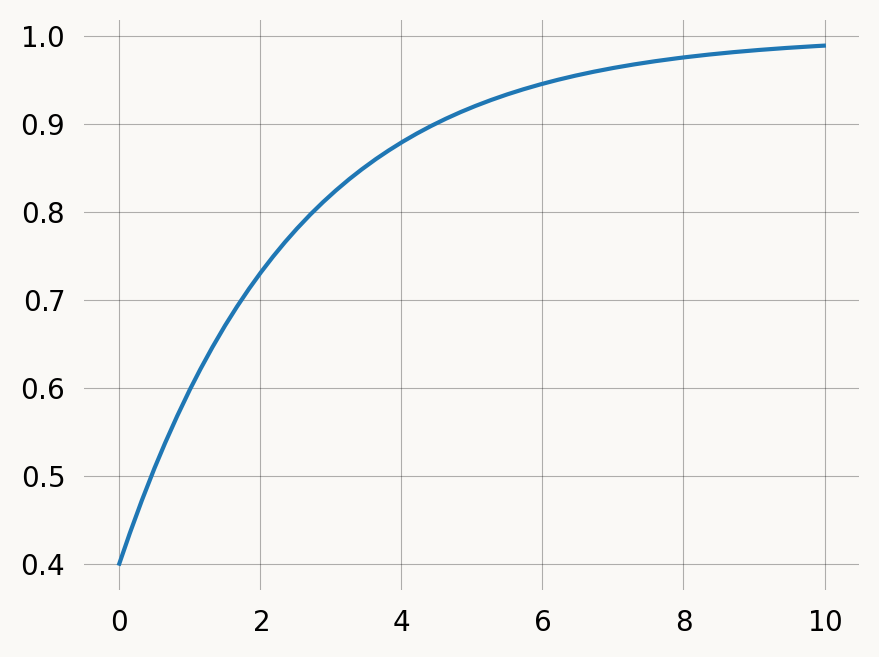
plt.show()You can treat the discrete graph by passing discrete=False (omit the argument), in which case the returned distribution is over continuous time, rather than discrete jumps:
rate = 0.4
graph = exponential(rate)
rewards = graph.discretize(1-rate)
x = np.arange(0, 10, 0.01)
# cdf = graph.reward_transform(rewards).cdf(x, discrete=False)
cdf = graph.reward_transform(rewards).cdf(x)
plt.plot(x, cdf)
plt.show()To model discrete mutations with rate 0.1 along branches of the coalescent tree. We could of course just make a new state-space creation function, but we can also manipulate an existing continuous one.
E.g. if you want to get the distribution of accumulated mutations over time rather than jumps.
Let’s model discrete mutations with rate 0.1 along branches of the coalescent tree. We could of course just make a new state-space creation function, but we can also manipulate an existing continuous one.
nr_samples = 4
@with_ipv([nr_samples]+[0]*(nr_samples-1))
def coalescent(state):
transitions = []
for i in range(state.size):
for j in range(i, state.size):
same = int(i == j)
if same and state[i] < 2:
continue
if not same and (state[i] < 1 or state[j] < 1):
continue
new = state.copy()
new[i] -= 1
new[j] -= 1
new[i+j+1] += 1
transitions.append((new, state[i]*(state[j]-same)/(1+same)))
return transitions
mutation_graph = Graph(coalescent)
mutation_graph.plot()This process is automated by the discretize method, which accepts either a single rate of discrete events for all state (E.g. graph.discretize(0.7)), or a callable mapping each state vector to a rate. The following example produces the mutation graph above:
from typing import Optional
mutation_graph = Graph(coalescent)
def mutation(state:np.ndarray, mutation_rate:float):
nr_lineages = sum(state)
return nr_lineages * mutation_rate
mutation_rate = 0.1
rewards = mutation_graph.discretize(mutation, mutation_rate=mutation_rate)The auxiliary nodes added have dummy states with all zeros and are labelled as “AUX” vertices when the graph is plotted:
If you want, you can assign nodes to subgraphs either by index or by state, by passing a callable as the by_index or by_state argument. For example, we can group the auxiliary nodes we added above into a separate subgraph:
def fun(index):
if rewards[index]:
return "Auxiliary states\ncollecting a\nreward (mutations)\neach time visited"
else:
return "Standard states"
mutation_graph.plot(by_index=fun)To find compute the expected number of mutations, we apply rewards such that the mutation vertex get a reward of 1 and all other states get reward of 0. Since this is already encoded in the mutation_vertices array we made above, we can convert it to integers and pass it as rewards:
mutation_graph.expectation(rewards)0.3666666666666667We can verify that the expected number of mutations correspond to the total branch length scaled with the mutation rate:
graph = Graph(coalescent)
tot_brlen = graph.expectation(graph.states().T.sum(axis=0))
tot_brlen * mutation_rate0.36666666666666664Or compare it to emprical mean of samples from the models
samples = mutation_graph.sample(10000, rewards=rewards)
samples.mean().item()0.3749samples = graph.sample(10000, rewards=graph.states().T.sum(axis=0))
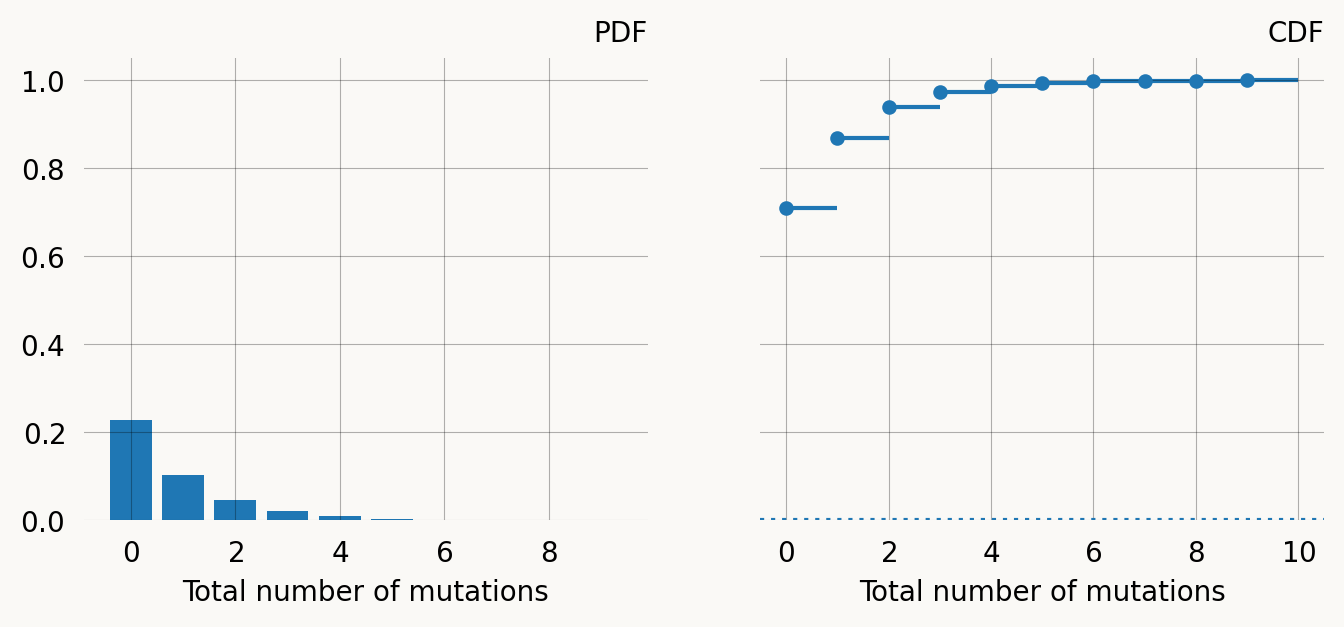
samples.mean().item() * mutation_rate0.3679581289515518If we reward transform the graph, we can find the distribution function for the the total number of mutations:
rev_trans_mutation_graph = mutation_graph.reward_transform_discrete(rewards)
rev_trans_mutation_graph.plot()Notice that with this reward transformation the graph is no longer sparse, as all paths through the graph are represented.
mutations = np.arange(10)
pmf = rev_trans_mutation_graph.pdf(mutations)
cdf = rev_trans_mutation_graph.cdf(mutations)
fig, (ax1, ax2) = plt.subplots(1, 2, figsize=(8, 3), sharey=True)
x = np.arange(0, 10, 1)
ax1.bar(x, pmf)
ax1.set_title("PDF")
ax1.set_xlabel('Total number of mutations')
left, right = x, np.arange(1, x.size+1, 1)
ax2.hlines(y=cdf, xmin=left, xmax=right, zorder=1)
ax2.scatter(left, cdf, s=18, zorder=2)
ax2.set_title("CDF")
ax2.set_xlabel('Total number of mutations')
ax2.axhline(y=mutation_graph.defect(), linestyle='dotted') ;Multivariate phase-type distributions (marginal)
We can compute marginal expectations using rewards. Instead of a univariate reward, we can have the distribution earn a vector of rewards for every time unit spent at each vertex. We can use the columns in our state matrix as reward vectors. Singleton rewards are:
graph = Graph(coalescent)
reward_matrix = graph.states().T
singleton = reward_matrix[0]
doubleton = reward_matrix[1]
tripleton = reward_matrix[2]
print(f"Singleton rewards: {singleton}")Singleton rewards: [0 4 2 0 1 0]Expected singleton branch length:
graph.expectation(singleton)1.9999999999999996Covariance of doubleton and tripleton branch length:
graph.covariance(singleton, doubleton)-0.22222222222222232If need be, we can also sample from the multivariate distribution:
nsims = 1000000
samples = graph.sample_multivariate(nsims, graph.states())
samples.shape(4, 1000000)Sanity check: The emprical covariance is ok:
float(sum(samples[0,:]*samples[1,:])/nsims - sum(samples[0,:])/nsims*sum(samples[1,:])/nsims)-0.21739365791458232