from phasic import (
Graph, with_ipv,
# Priors
GaussPrior, HalfCauchyPrior, DataPrior,
# Step size schedules
ConstantStepSize, ExpStepSize, AdaptiveStepSize, WarmupExpStepSize,
# Regularization schedules
ConstantRegularization, ExpRegularization, ExponentialCDFRegularization,
# Optimizers (for examples)
Adam,
) # ALWAYS import phasic first to set jax backend correctly
import numpy as np
import jax.numpy as jnp
import matplotlib.pyplot as plt
import seaborn as sns
np.random.seed(42)
try:
from vscodenb import set_vscode_theme
set_vscode_theme()
except ImportError:
pass
sns.set_palette('tab10')Priors and schedules
This page covers the prior distributions, step size schedules, and regularization schedules available for use with SVGD inference.
Priors
GaussPrior
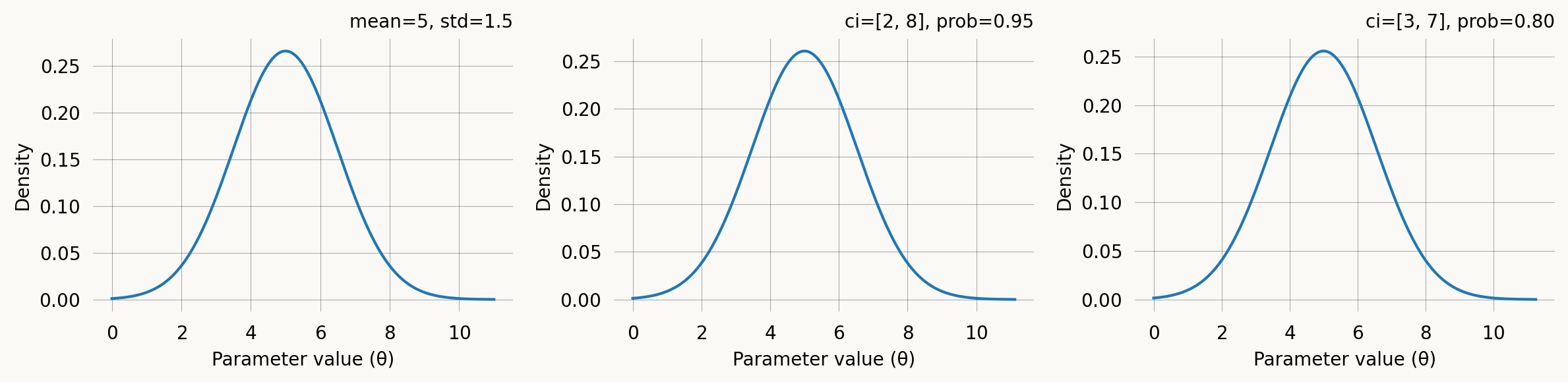
A Gaussian (normal) prior centred on a point estimate with a given spread. It can be specified either by the mean and standard deviation or by a credible interval. E.g., “I believe the parameter is between 2 and 8 with 95% probability” translates directly to GaussPrior(ci=(2, 8)). The two specifications are equivalent: a 95% credible interval [low, high] maps to mean = (low + high) / 2 and std = (high - low) / (2 * z_0.975).
fig, axes = plt.subplots(1, 3, figsize=(12, 3))
# 1. Mean + std
p1 = GaussPrior(mean=5.0, std=1.5)
p1.plot(ax=axes[0], return_ax=True)
axes[0].set_title(f'mean=5, std=1.5')
# 2. 95% credible interval
p2 = GaussPrior(ci=(2.0, 8.0))
p2.plot(ax=axes[1], return_ax=True)
axes[1].set_title(f'ci=[2, 8], prob=0.95')
# 3. 80% credible interval (tighter)
p3 = GaussPrior(ci=(3.0, 7.0), prob=0.80)
p3.plot(ax=axes[2], return_ax=True)
axes[2].set_title(f'ci=[3, 7], prob=0.80')
plt.tight_layout()HalfCauchyPrior
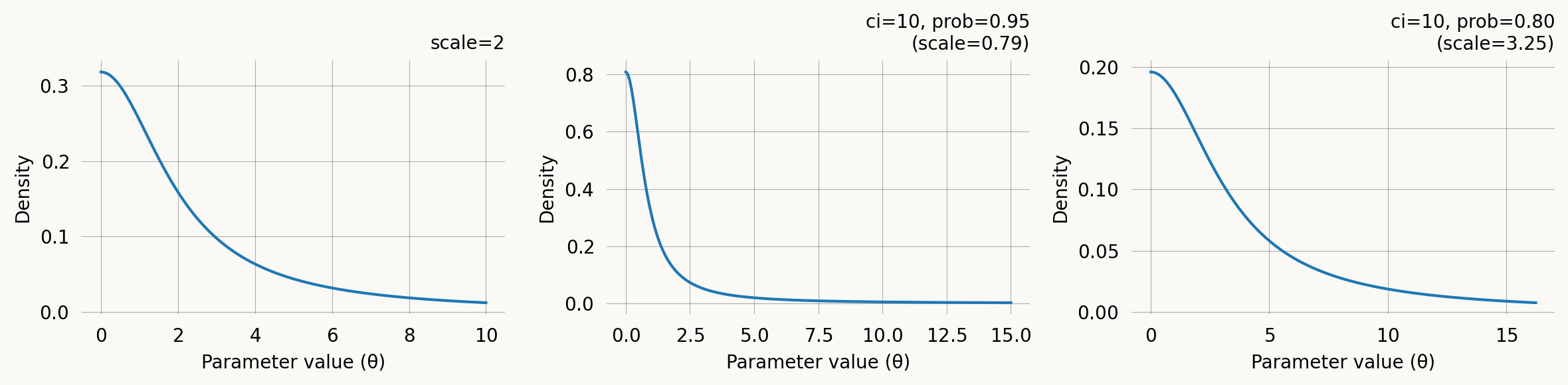
A half-Cauchy prior with support on (0, \infty). Its heavy tails make it a popular weakly informative prior for scale and rate parameters. Concentrates mass near zero but allows for large values.
f(\theta) = \frac{2}{\pi \, \gamma \left(1 + (\theta/\gamma)^2\right)}, \quad \theta > 0
It can be specified via the scale parameter \gamma directly, or via a credible-interval upper bound. The CI specification says: “I believe there is a prob probability that the parameter is below ci.” A lower prob for the same ci implies a larger scale (heavier tail).
fig, axes = plt.subplots(1, 3, figsize=(12, 3))
# 1. Scale parameter
h1 = HalfCauchyPrior(scale=2.0)
h1.plot(ax=axes[0], return_ax=True)
axes[0].set_title('scale=2')
# 2. 95% CI upper bound
h2 = HalfCauchyPrior(ci=10.0)
h2.plot(ax=axes[1], return_ax=True)
axes[1].set_title(f'ci=10, prob=0.95\n(scale={h2.scale:.2f})')
# 3. 80% CI upper bound
h3 = HalfCauchyPrior(ci=10.0, prob=0.80)
h3.plot(ax=axes[2], return_ax=True)
axes[2].set_title(f'ci=10, prob=0.80\n(scale={h3.scale:.2f})')
plt.tight_layout()DataPrior
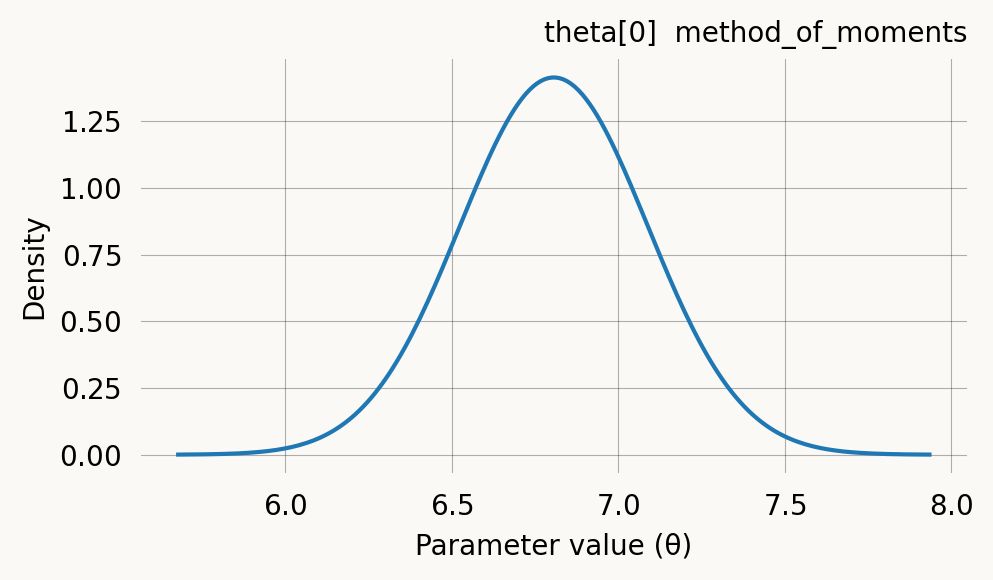
DataPrior constructs a data-informed prior automatically. Instead of specifying mean and spread by hand, it estimates them from the observed data using either method of moments (standard phase-type graphs) or probability matching (joint probability graphs). The graph type is detected automatically. The result is a list of per-parameter GaussPrior objects (with None at fixed-parameter positions) that integrates directly with graph.svgd().
# Build a simple coalescent model
nr_samples = 4
@with_ipv([nr_samples]+[0]*(nr_samples-1))
def coalescent_1param(state):
transitions = []
for i in range(state.size):
for j in range(i, state.size):
same = int(i == j)
if same and state[i] < 2:
continue
if not same and (state[i] < 1 or state[j] < 1):
continue
new = state.copy()
new[i] -= 1
new[j] -= 1
new[i+j+1] += 1
transitions.append([new, [state[i]*(state[j]-same)/(1+same)]])
return transitions
graph = Graph(coalescent_1param)
true_theta = [7.0]
graph.update_weights(true_theta)
data = graph.sample(1000)dp = DataPrior(graph, data, sd=2.0)
print(dp)
print(f"Method: {dp.method}")
print(f"Theta: {dp.theta}")
print(f"Std: {dp.std}")
print(f"Converged: {dp.success}")DataPrior(method=method_of_moments, converged, theta=[6.8063])
Method: method_of_moments
Theta: [6.80628673]
Std: [0.14097399]
Converged: TrueDataPrior is iterable — you can inspect individual per-parameter priors and pass it directly to graph.svgd() as the prior argument:
# Inspect per-parameter priors
print(f"Number of parameters: {len(dp)}")
for i, p in enumerate(dp):
if p is not None:
print(f" theta[{i}]: GaussPrior(mean={p.mu:.3f}, std={p.sigma:.3f})")
else:
print(f" theta[{i}]: None (fixed)")Number of parameters: 1
theta[0]: GaussPrior(mean=6.806, std=0.282)# Use directly with SVGD
svgd = graph.svgd(data, prior=dp, optimizer=Adam(0.25))
svgd.summary()Parameter Fixed MAP Mean SD HPD 95% lo HPD 95% hi
0 No 6.7121 6.7367 0.1025 6.5919 6.9032
Particles: 24, Iterations: 100See the API documentation on DataPrior for all parameters.
Per-parameter priors
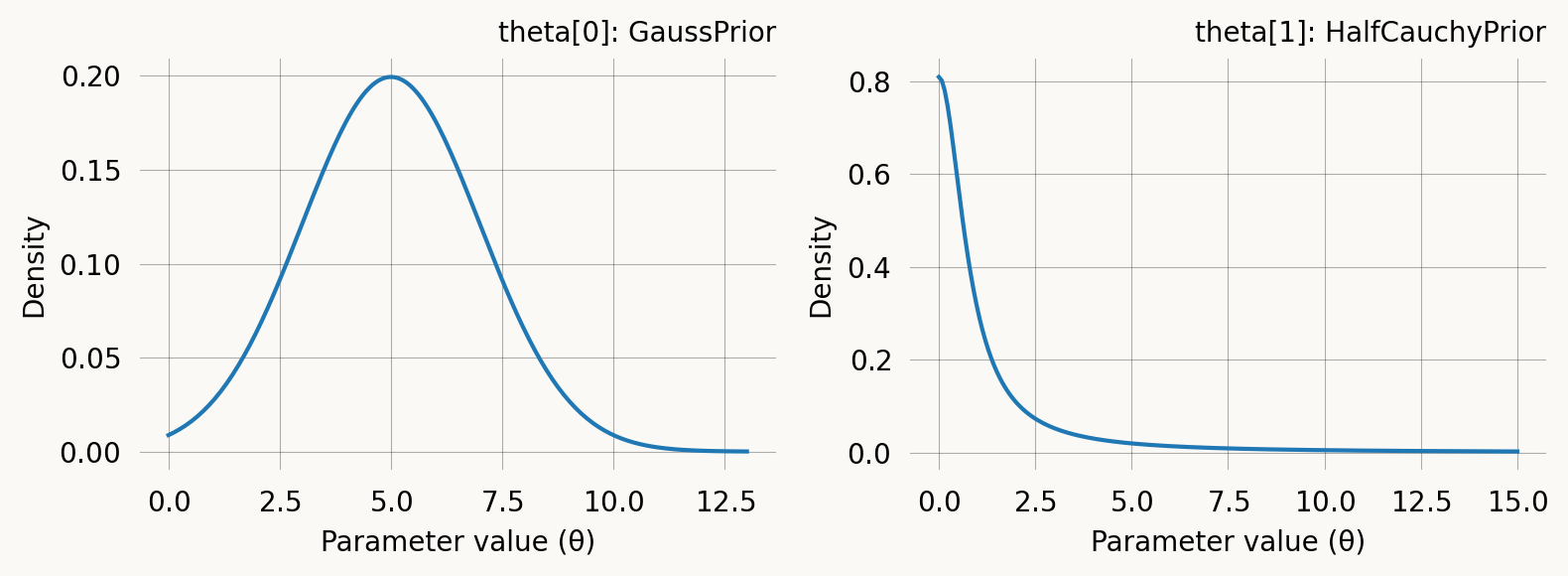
When a model has multiple parameters, you can assign a different prior to each one by passing a list of prior objects to graph.svgd(). Use None at positions corresponding to fixed parameters:
per_param_priors = [
GaussPrior(mean=5.0, std=2.0), # theta[0]
HalfCauchyPrior(ci=10.0), # theta[1]
]
fig, axes = plt.subplots(1, 2, figsize=(8, 3))
per_param_priors[0].plot(ax=axes[0], return_ax=True)
axes[0].set_title('theta[0]: GaussPrior')
per_param_priors[1].plot(ax=axes[1], return_ax=True)
axes[1].set_title('theta[1]: HalfCauchyPrior')
plt.tight_layout()Step size schedules
The step size (learning rate) controls how far particles move at each SVGD iteration. A step size that is too large causes oscillation; too small causes slow convergence. Schedules let you vary the step size over the course of optimization. Pass a schedule to graph.svgd() via the learning_rate parameter, or to an optimizer like Adam(learning_rate=schedule).
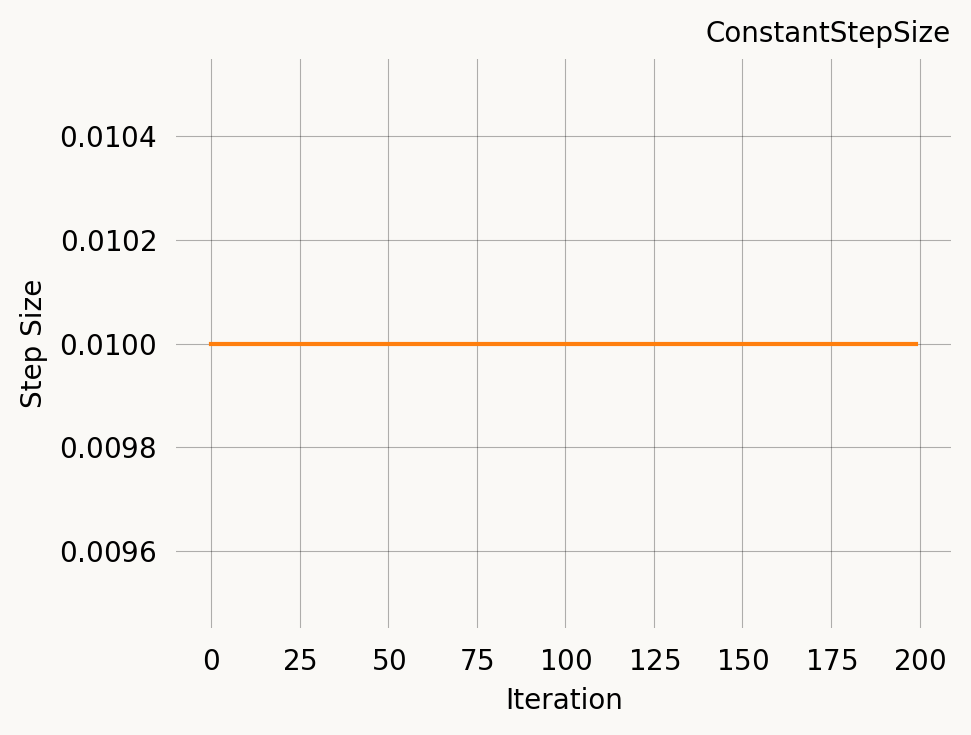
ConstantStepSize
The simplest schedule: the same step size at every iteration. This is also the implicit default when you pass a plain number as the learning_rate.
const = ConstantStepSize(0.01)
print(f"Step at iteration 0: {const(0)}")
print(f"Step at iteration 500: {const(500)}")
const.plot(200)Step at iteration 0: 0.01
Step at iteration 500: 0.01Se the API documentation for ConstantStepSize for all parameters.
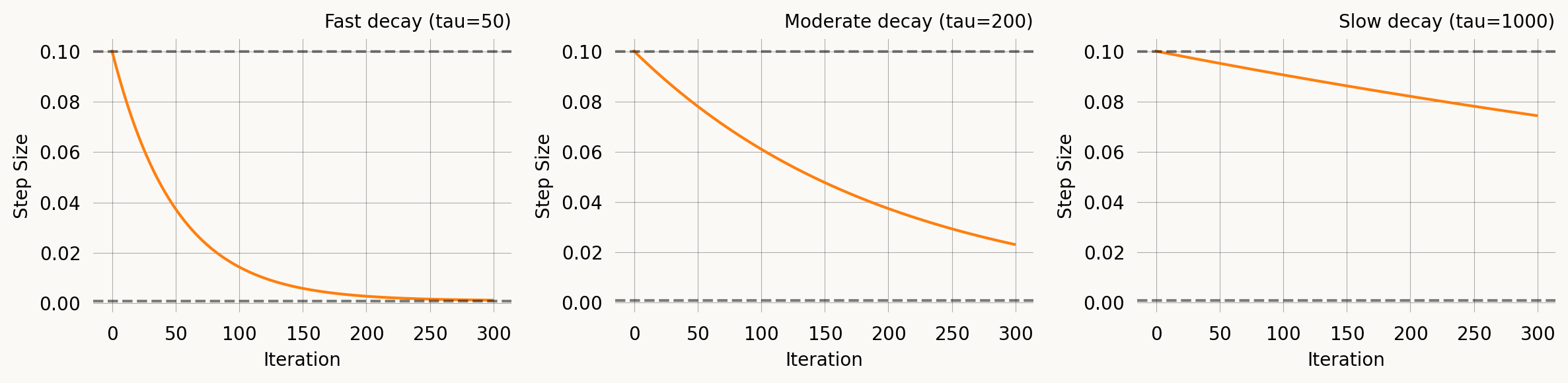
ExpStepSize
Exponential decay from first_step to last_step with time constant tau:
\eta(t) = \eta_0 \, e^{-t/\tau} + \eta_\infty \left(1 - e^{-t/\tau}\right)
At t = \tau, roughly 63% of the decay has occurred. This is the most commonly used schedule — start with a large step to explore, then settle down for fine-tuning.
fig, axes = plt.subplots(1, 3, figsize=(12, 3))
# Fast decay
s1 = ExpStepSize(first_step=0.1, last_step=0.001, tau=50.0)
s1.plot(300, ax=axes[0], return_ax=True)
axes[0].set_title('Fast decay (tau=50)')
# Moderate decay
s2 = ExpStepSize(first_step=0.1, last_step=0.001, tau=200.0)
s2.plot(300, ax=axes[1], return_ax=True)
axes[1].set_title('Moderate decay (tau=200)')
# Slow decay
s3 = ExpStepSize(first_step=0.1, last_step=0.001, tau=1000.0)
s3.plot(300, ax=axes[2], return_ax=True)
axes[2].set_title('Slow decay (tau=1000)')
plt.tight_layout()Se the API documentation for ExpStepSize for all parameters.
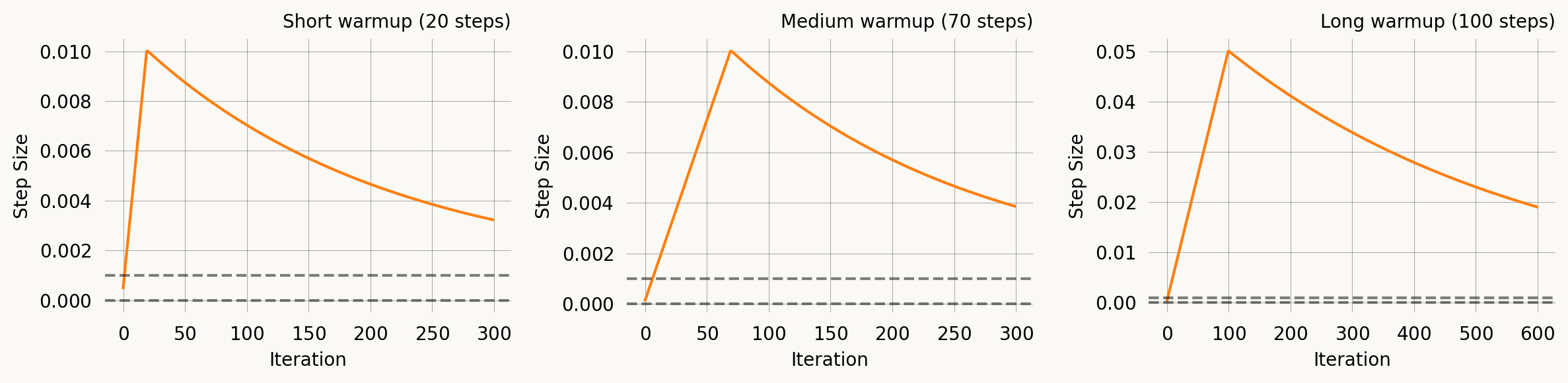
WarmupExpStepSize
A linear ramp-up phase followed by exponential decay. This is useful with Adam and other adaptive optimizers where moment estimates are poorly calibrated in the first few iterations — a warmup prevents large early updates.
\eta(t) = \begin{cases} \eta_{\text{peak}} \cdot \frac{t+1}{T_{\text{warmup}}} & t < T_{\text{warmup}} \\ \eta_{\text{peak}} \, e^{-(t - T_{\text{warmup}})/\tau} + \eta_\infty \left(1 - e^{-(t - T_{\text{warmup}})/\tau}\right) & t \geq T_{\text{warmup}} \end{cases}
fig, axes = plt.subplots(1, 3, figsize=(12, 3))
# Short warmup
w1 = WarmupExpStepSize(peak_lr=0.01, warmup_steps=20, last_lr=0.001, tau=200.0)
w1.plot(300, ax=axes[0], return_ax=True)
axes[0].set_title('Short warmup (20 steps)')
# Medium warmup
w2 = WarmupExpStepSize(peak_lr=0.01, warmup_steps=70, last_lr=0.001, tau=200.0)
w2.plot(300, ax=axes[1], return_ax=True)
axes[1].set_title('Medium warmup (70 steps)')
# Long warmup, high peak
w3 = WarmupExpStepSize(peak_lr=0.05, warmup_steps=100, last_lr=0.001, tau=500.0)
w3.plot(600, ax=axes[2], return_ax=True)
axes[2].set_title('Long warmup (100 steps)')
plt.tight_layout()Se the API documentation for WarmupExpStepSize for all parameters.
AdaptiveStepSize
Adjusts the step size dynamically based on particle spread. When particles are too concentrated the step size increases; when too dispersed it decreases. This is a simple heuristic that does not require tuning a decay schedule, but its behaviour depends on the optimisation trajectory.
adaptive = AdaptiveStepSize(base_step=0.01, kl_target=0.1, adjust_rate=0.1)
print(f"Base step: {adaptive.base_step}")
print(f"Without particles: {adaptive(0):.4f}")
# Simulate concentrated particles
concentrated = jnp.array([[1.0], [1.01], [0.99]])
print(f"Concentrated particles: {adaptive(1, concentrated):.4f} (increases)")
# Simulate spread particles
spread = jnp.array([[1.0], [10.0], [100.0]])
print(f"Spread particles: {adaptive(2, spread):.4f} (decreases)")Base step: 0.01
Without particles: 0.0100
Concentrated particles: 0.0110 (increases)
Spread particles: 0.0099 (decreases)Se the API documentation for AdaptiveStepSize for all parameters.
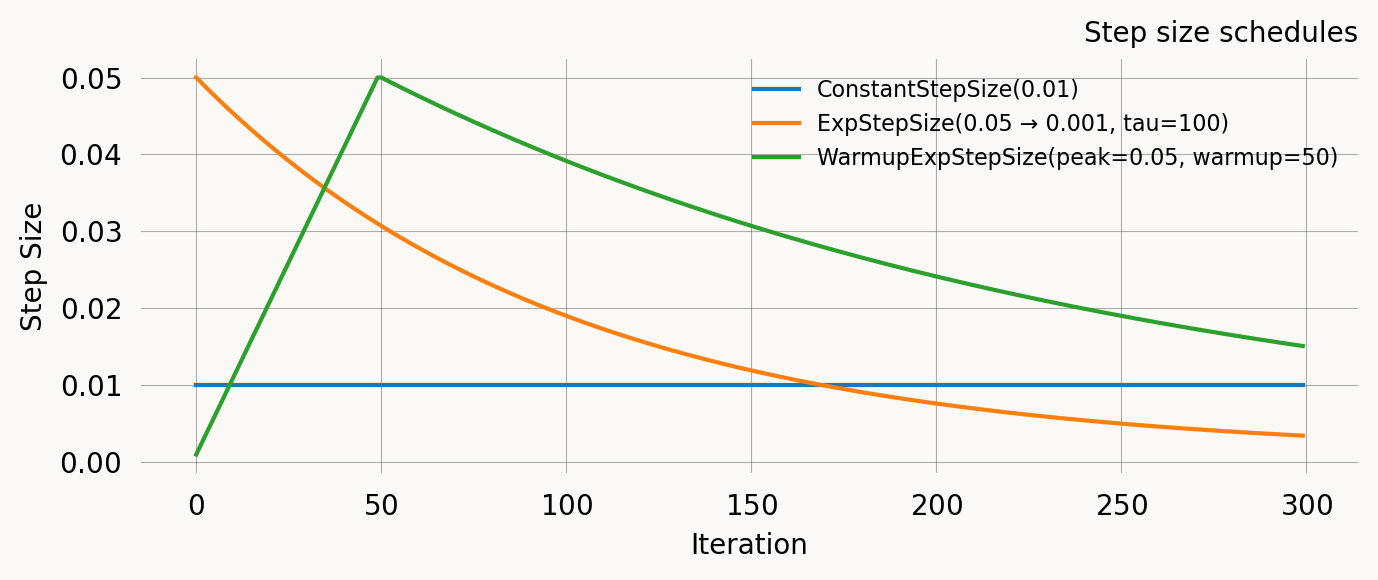
Comparison
All four schedules on the same axes for a 300-iteration run:
nr_iter = 300
iters = np.arange(nr_iter)
schedules = {
'ConstantStepSize(0.01)': ConstantStepSize(0.01),
'ExpStepSize(0.05 → 0.001, tau=100)': ExpStepSize(0.05, 0.001, 100.0),
'WarmupExpStepSize(peak=0.05, warmup=50)': WarmupExpStepSize(0.05, 50, 0.001, 200.0),
}
fig, ax = plt.subplots(figsize=(7, 3))
for label, sched in schedules.items():
vals = [float(sched(i)) for i in iters]
ax.plot(iters, vals, label=label)
ax.set_xlabel('Iteration')
ax.set_ylabel('Step Size')
ax.set_title('Step size schedules')
ax.legend(fontsize=8)
plt.tight_layout()Regularization schedules
Moment regularization adds a penalty term to the SVGD objective that encourages the model moments at the current parameter values to match the empirical moments. This is controlled by the regularization parameter in graph.svgd() and requires specifying nr_moments. Strong regularization early in optimization helps guide particles into a reasonable region; reducing it later lets the likelihood dominate for fine-tuning.
ConstantRegularization
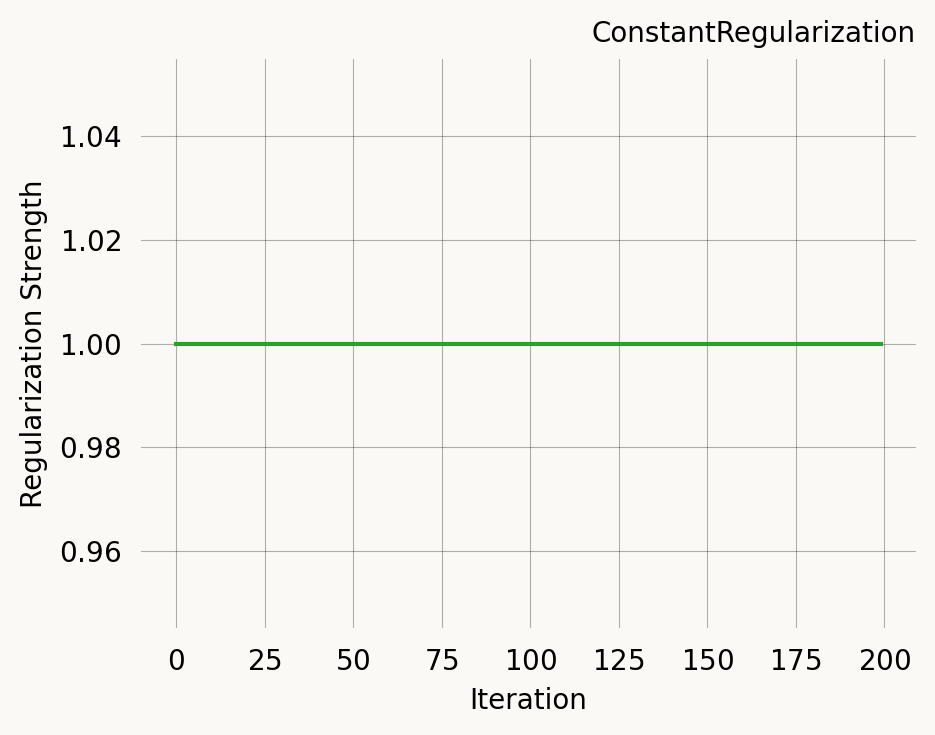
A fixed regularization strength throughout optimisation.
const_reg = ConstantRegularization(1.0)
print(f"Strength at iteration 0: {const_reg(0)}")
print(f"Strength at iteration 500: {const_reg(500)}")
const_reg.plot(200)Strength at iteration 0: 1.0
Strength at iteration 500: 1.0Se the API documentation for ConstantRegularization for all parameters.
ExpRegularization
Exponential decay, identical in form to ExpStepSize:
\lambda(t) = \lambda_0 \, e^{-t/\tau} + \lambda_\infty \left(1 - e^{-t/\tau}\right)
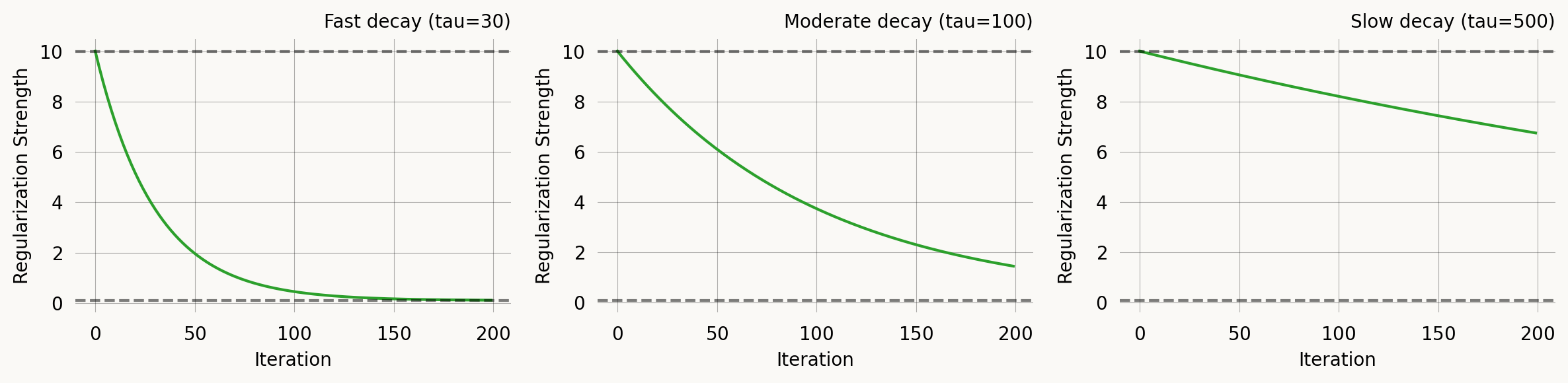
Start with strong regularization to guide particles, then reduce to let the likelihood dominate.
fig, axes = plt.subplots(1, 3, figsize=(12, 3))
r1 = ExpRegularization(first_reg=10.0, last_reg=0.1, tau=30.0)
r1.plot(200, ax=axes[0], return_ax=True)
axes[0].set_title('Fast decay (tau=30)')
r2 = ExpRegularization(first_reg=10.0, last_reg=0.1, tau=100.0)
r2.plot(200, ax=axes[1], return_ax=True)
axes[1].set_title('Moderate decay (tau=100)')
r3 = ExpRegularization(first_reg=10.0, last_reg=0.1, tau=500.0)
r3.plot(200, ax=axes[2], return_ax=True)
axes[2].set_title('Slow decay (tau=500)')
plt.tight_layout()Se the API documentation for ExpRegularization for all parameters.
ExponentialCDFRegularization
Uses the exponential CDF for a smooth S-curve transition:
\lambda(t) = \lambda_0 + (\lambda_\infty - \lambda_0) \left(1 - e^{-t/\tau}\right)
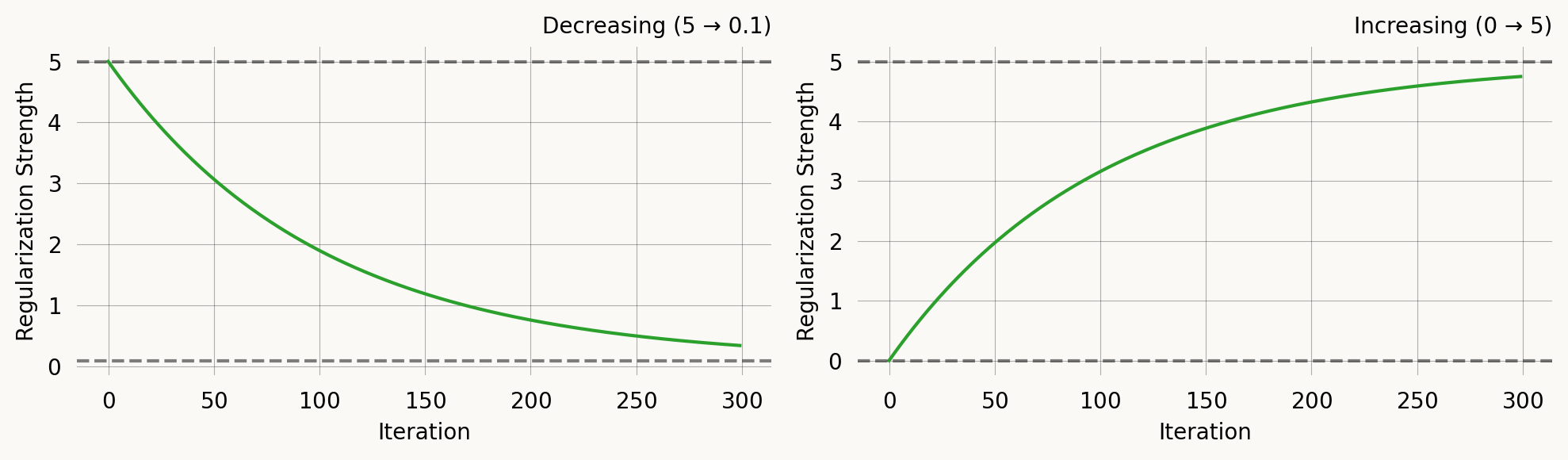
This is mathematically equivalent to ExpRegularization for decreasing schedules, but the parameterisation makes it equally natural for increasing schedules — useful for progressive regularization where you start with pure likelihood and gradually add moment matching.
fig, axes = plt.subplots(1, 2, figsize=(10, 3))
# Decreasing (same as ExpRegularization)
c1 = ExponentialCDFRegularization(first_reg=5.0, last_reg=0.1, tau=100.0)
c1.plot(300, ax=axes[0], return_ax=True)
axes[0].set_title('Decreasing (5 → 0.1)')
# Increasing — start with likelihood only, add regularization later
c2 = ExponentialCDFRegularization(first_reg=0.0, last_reg=5.0, tau=100.0)
c2.plot(300, ax=axes[1], return_ax=True)
axes[1].set_title('Increasing (0 → 5)')
plt.tight_layout()Se the API documentation for ExponentialCDFRegularization for all parameters.
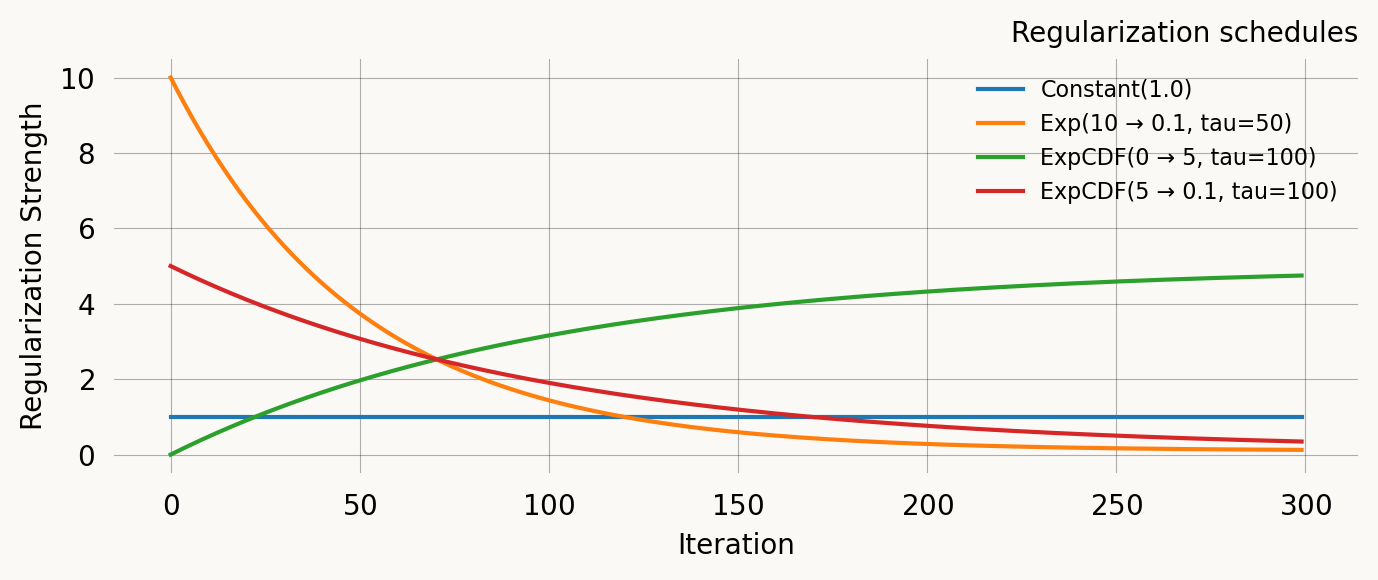
Comparison
nr_iter = 300
iters = np.arange(nr_iter)
reg_schedules = {
'Constant(1.0)': ConstantRegularization(1.0),
'Exp(10 → 0.1, tau=50)': ExpRegularization(10.0, 0.1, 50.0),
'ExpCDF(0 → 5, tau=100)': ExponentialCDFRegularization(0.0, 5.0, 100.0),
'ExpCDF(5 → 0.1, tau=100)': ExponentialCDFRegularization(5.0, 0.1, 100.0),
}
fig, ax = plt.subplots(figsize=(7, 3))
for label, sched in reg_schedules.items():
vals = [float(sched(i)) for i in iters]
ax.plot(iters, vals, label=label)
ax.set_xlabel('Iteration')
ax.set_ylabel('Regularization Strength')
ax.set_title('Regularization schedules')
ax.legend(fontsize=8)
plt.tight_layout()Putting it together
A typical SVGD call combining priors and schedules:
svgd = graph.svgd(
data,
prior=DataPrior(graph, data, sd=2.0),
optimizer=Adam(
learning_rate=ExpStepSize(first_step=0.1, last_step=0.01, tau=30.0),
),
# regularization=ExpRegularization(first_reg=5.0, last_reg=0.1, tau=20.0),
# nr_moments=2,
n_iterations=200,
)
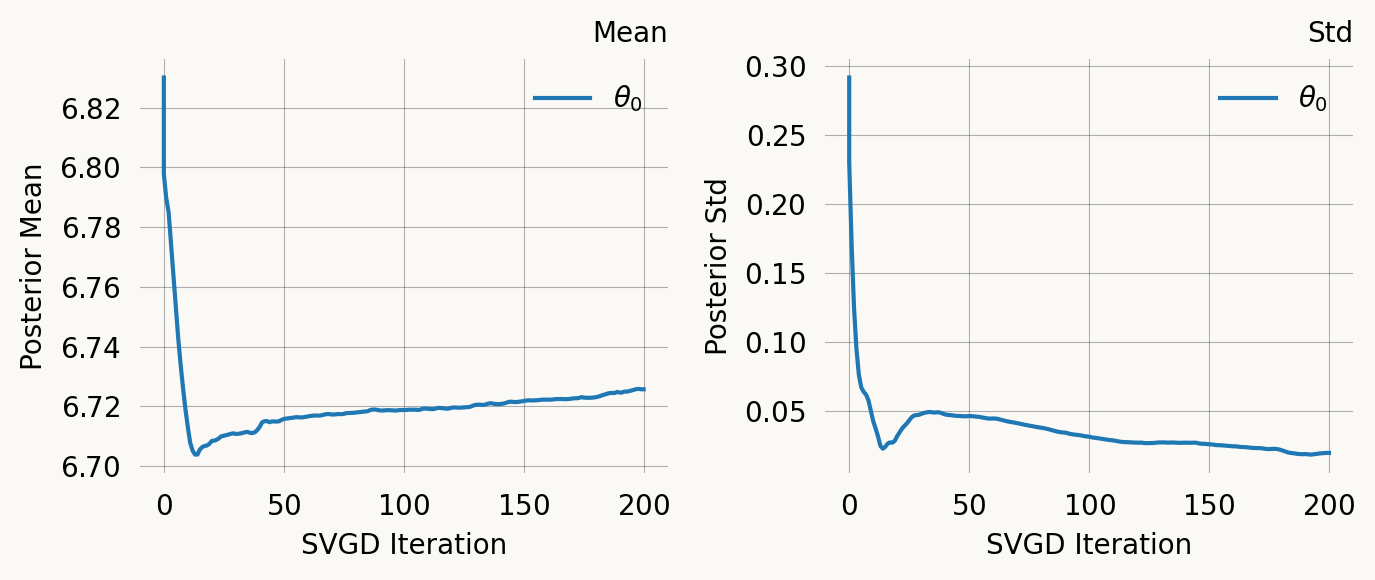
svgd.summary()
svgd.plot_convergence() ;Parameter Fixed MAP Mean SD HPD 95% lo HPD 95% hi
0 No 6.7210 6.7257 0.0198 6.6942 6.7676
Particles: 24, Iterations: 200